Characterization of the First Virulent Phage Infecting Oenococcus oeni, the Queen of the Cellars
- PMID: 33519734
- PMCID: PMC7838156
- DOI: 10.3389/fmicb.2020.596541
Characterization of the First Virulent Phage Infecting Oenococcus oeni, the Queen of the Cellars
Abstract
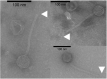
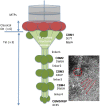
There has been little exploration of how phages contribute to the diversity of the bacterial community associated with winemaking and may impact fermentations and product quality. Prophages of Oenococcus oeni, the most common species of lactic acid bacteria (LAB) associated with malolactic fermentation of wine, have been described, but no data is available regarding phages of O. oeni with true virulent lifestyles. The current study reports on the incidence and characterization of the first group of virulent oenophages named Vinitor, isolated from the enological environment. Vinitor phages are morphologically very similar to siphoviruses infecting other LAB. Although widespread during winemaking, they are more abundant in musts than temperate oenophages. We obtained the complete genomic sequences of phages Vinitor162 and Vinitor27, isolated from white and red wines, respectively. The assembled genomes shared 97.6% nucleotide identity and belong to the same species. Coupled with phylogenetic analysis, our study revealed that the genomes of Vinitor phages are architecturally mosaics and represent unique combinations of modules amongst LAB infecting-phages. Our data also provide some clues to possible evolutionary connections between Vinitor and (pro)phages associated to epiphytic and insect-related bacteria.
Keywords: Oenococcus; fermentation; grapes; lactic acid bacteria; lytic phage; phage (bacteriophage); phage phylogenetics; wine.
Copyright © 2021 Philippe, Chaïb, Jaomanjaka, Claisse, Lucas, Samot, Cambillau and Le Marrec.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Exploring the diversity of bacteriophage specific to Oenococcus oeni and Lactobacillus spp and their role in wine production.Appl Microbiol Biotechnol. 2021 Dec;105(23):8575-8592. doi: 10.1007/s00253-021-11509-2. Epub 2021 Oct 25. Appl Microbiol Biotechnol. 2021. PMID: 34694447 Review.
-
Characterization of strictly lytic phages infecting Oenococcus oeni from Merlot wines and proposal of a new genus.Microbiol Spectr. 2025 Aug 12:e0258824. doi: 10.1128/spectrum.02588-24. Online ahead of print. Microbiol Spectr. 2025. PMID: 40792506
-
Phage-host interactions analysis of newly characterized Oenococcus oeni bacteriophages: Implications for malolactic fermentation in wine.Int J Food Microbiol. 2017 Apr 4;246:12-19. doi: 10.1016/j.ijfoodmicro.2017.01.020. Epub 2017 Jan 31. Int J Food Microbiol. 2017. PMID: 28189899
-
Lysogeny in the Lactic Acid Bacterium Oenococcus oeni Is Responsible for Modified Colony Morphology on Red Grape Juice Agar.Appl Environ Microbiol. 2019 Sep 17;85(19):e00997-19. doi: 10.1128/AEM.00997-19. Print 2019 Oct 1. Appl Environ Microbiol. 2019. PMID: 31375489 Free PMC article.
-
Distribution of Oenococcus oeni populations in natural habitats.Appl Microbiol Biotechnol. 2019 Apr;103(7):2937-2945. doi: 10.1007/s00253-019-09689-z. Epub 2019 Feb 20. Appl Microbiol Biotechnol. 2019. PMID: 30788540 Free PMC article. Review.
Cited by
-
Investigation of the Phageome and Prophages in French Cider, a Fermented Beverage.Microorganisms. 2022 Jun 12;10(6):1203. doi: 10.3390/microorganisms10061203. Microorganisms. 2022. PMID: 35744720 Free PMC article.
-
Exploring the diversity of bacteriophage specific to Oenococcus oeni and Lactobacillus spp and their role in wine production.Appl Microbiol Biotechnol. 2021 Dec;105(23):8575-8592. doi: 10.1007/s00253-021-11509-2. Epub 2021 Oct 25. Appl Microbiol Biotechnol. 2021. PMID: 34694447 Review.
-
Characterization of Parageobacillus Bacteriophage vB_PtoS_NIIg3.2-A Representative of a New Genus within Thermophilic Siphoviruses.Int J Mol Sci. 2023 Sep 12;24(18):13980. doi: 10.3390/ijms241813980. Int J Mol Sci. 2023. PMID: 37762288 Free PMC article.
-
Wine Fermentation as a Model System for Microbial Ecology and Evolution.Environ Microbiol. 2025 Apr;27(4):e70092. doi: 10.1111/1462-2920.70092. Environ Microbiol. 2025. PMID: 40222749 Free PMC article. Review.
-
AlphaFold2 in biomedical research: facilitating the development of diagnostic strategies for disease.Front Mol Biosci. 2024 Jul 30;11:1414916. doi: 10.3389/fmolb.2024.1414916. eCollection 2024. Front Mol Biosci. 2024. PMID: 39139810 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

