Comparative Transcriptome Profile Analysis of Longissimus dorsi Muscle Tissues From Two Goat Breeds With Different Meat Production Performance Using RNA-Seq
- PMID: 33519920
- PMCID: PMC7838615
- DOI: 10.3389/fgene.2020.619399
Comparative Transcriptome Profile Analysis of Longissimus dorsi Muscle Tissues From Two Goat Breeds With Different Meat Production Performance Using RNA-Seq
Abstract
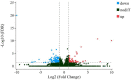
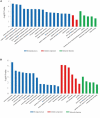
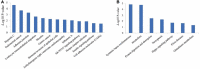
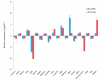
Carcass weight, meat quality and muscle components are important traits economically and they underpin most of the commercial return to goat producers. In this study, the Longissimus dorsi muscle tissues were collected from five Liaoning cashmere (LC) goats and five Ziwuling black (ZB) goats with phenotypic difference in carcass weight, some meat quality traits and muscle components. The histological quantitative of collagen fibers and the transcriptome profiles in the Longissimus dorsi muscle tissues were investigated using Masson-trichrome staining and RNA-Seq, respectively. The percentage of total collagen fibers in the Longissimus dorsi muscle tissues from ZB goats was less than those from LC goats, suggesting that these ZB goats had more tender meat. An average of 15,919 and 15,582 genes were found to be expressed in Longissimus dorsi muscle tissues from LC and ZB goats, respectively. Compared to LC goats, the expression levels of 78 genes were up-regulated in ZB goats, while 133 genes were down-regulated. Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses revealed that the differentially expressed genes (DEGs) were significantly enriched in GO terms related to the muscle growth and development and the deposition of intramuscular fat and lipid metabolism, hippo signaling pathway and Jak-STAT signaling pathway. The results provide an improved understanding of the genetic mechanisms regulating meat production performance in goats, and will help us improve the accuracy of selection for meat traits in goats using marker-assisted selection based on these differentially expressed genes obtained.
Keywords: Liaoning cashmere goats; RNA-Seq; Ziwuling black goats; differentially expressed gene; skeletal muscle.
Copyright © 2021 Shen, Hao, Wang, Hu, Liu, Li, Ke, Song, Lu, Hu, Qiao, Wu and Luo.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
-
- Albrecht E., Komolka K., Will K. (2013). Tissue distribution and blood circulation of agouti signaling protein (ASIP) in relationship with body composition in cattle. Proc. JPN. Soc. Anim. Nutr. Metab. 57 89–95.
-
- Arora R., Naveen Kumar S., Sudarshan S., Fairoze M. N., Kaur M., Sharma A., et al. (2019). Transcriptome profiling of longissimus thoracis muscles identifies highly connected differentially expressed genes in meat type sheep of India. PLoS One 14:e0217461. 10.1371/journal.pone.0217461 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

