Catalytic Mechanism of Non-Target DNA Cleavage in CRISPR-Cas9 Revealed by Ab Initio Molecular Dynamics
- PMID: 33520346
- PMCID: PMC7842700
- DOI: 10.1021/acscatal.0c03566
Catalytic Mechanism of Non-Target DNA Cleavage in CRISPR-Cas9 Revealed by Ab Initio Molecular Dynamics
Abstract
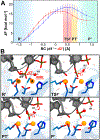
CRISPR-Cas9 is a cutting-edge genome editing technology, which uses the endonuclease Cas9 to introduce mutations at desired sites of the genome. This revolutionary tool is promising to treat a myriad of human genetic diseases. Nevertheless, the molecular basis of DNA cleavage, which is a fundamental step for genome editing, has not been established. Here, quantum-classical molecular dynamics (MD) and free energy methods are used to disclose the two-metal-dependent mechanism of phosphodiester bond cleavage in CRISPR-Cas9. Ab initio MD reveals a conformational rearrangement of the Mg2+-bound RuvC active site, which entails the relocation of H983 to act as a general base. Then, the DNA cleavage proceeds through a concerted associative pathway fundamentally assisted by the joint dynamics of the two Mg2+ ions. This clarifies previous controversial experimental evidence, which could not fully establish the catalytic role of the conserved H983 and the metal cluster conformation. The comparison with other two-metal-dependent enzymes supports the identified mechanism and suggests a common catalytic strategy for genome editing and recombination. Overall, the non-target DNA cleavage catalysis described here resolves a fundamental open question in the CRISPR-Cas9 biology and provides valuable insights for improving the catalytic efficiency and the metal-dependent function of the Cas9 enzyme, which are at the basis of the development of genome editing tools.
Keywords: CRISPR-Cas9; QM/MM; free energy simulations; genome editing; magnesium-aided catalysis; non-coding RNA; phosphodiester bond cleavage; protein/nucleic acid interactions.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
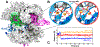


Similar articles
-
Principles of target DNA cleavage and the role of Mg2+ in the catalysis of CRISPR-Cas9.Nat Catal. 2022 Oct;5(10):912-922. doi: 10.1038/s41929-022-00848-6. Epub 2022 Oct 6. Nat Catal. 2022. PMID: 36778082 Free PMC article.
-
Structure and Dynamics of the CRISPR-Cas9 Catalytic Complex.J Chem Inf Model. 2019 May 28;59(5):2394-2406. doi: 10.1021/acs.jcim.8b00988. Epub 2019 Feb 27. J Chem Inf Model. 2019. PMID: 30763088
-
The Electronic Structure of Genome Editors from the First Principles.Electron Struct. 2023 Mar;5(1):014003. doi: 10.1088/2516-1075/acb410. Epub 2023 Feb 1. Electron Struct. 2023. PMID: 36926635 Free PMC article.
-
Insights into the Mechanism of CRISPR/Cas9-Based Genome Editing from Molecular Dynamics Simulations.ACS Omega. 2022 Dec 30;8(2):1817-1837. doi: 10.1021/acsomega.2c05583. eCollection 2023 Jan 17. ACS Omega. 2022. PMID: 36687047 Free PMC article. Review.
-
Potential pitfalls of CRISPR/Cas9-mediated genome editing.FEBS J. 2016 Apr;283(7):1218-31. doi: 10.1111/febs.13586. Epub 2015 Nov 27. FEBS J. 2016. PMID: 26535798 Review.
Cited by
-
Controlled Trafficking of Multiple and Diverse Cations Prompts Nucleic Acid Hydrolysis.ACS Catal. 2021 Jul 16;11(14):8786-8797. doi: 10.1021/acscatal.1c01825. Epub 2021 Jul 2. ACS Catal. 2021. PMID: 35145762 Free PMC article.
-
R-loop formation and conformational activation mechanisms of Cas9.Nature. 2022 Sep;609(7925):191-196. doi: 10.1038/s41586-022-05114-0. Epub 2022 Aug 24. Nature. 2022. PMID: 36002571 Free PMC article.
-
A Machine Learning Approach to Identify the Importance of Novel Features for CRISPR/Cas9 Activity Prediction.Biomolecules. 2022 Aug 16;12(8):1123. doi: 10.3390/biom12081123. Biomolecules. 2022. PMID: 36009017 Free PMC article.
-
Atomic-scale insights into allosteric inhibition and evolutional rescue mechanism of Streptococcus thermophilus Cas9 by the anti-CRISPR protein AcrIIA6.Comput Struct Biotechnol J. 2021 Nov 16;19:6108-6124. doi: 10.1016/j.csbj.2021.11.010. eCollection 2021. Comput Struct Biotechnol J. 2021. PMID: 34900128 Free PMC article.
-
Quantum biological insights into CRISPR-Cas9 sgRNA efficiency from explainable-AI driven feature engineering.Nucleic Acids Res. 2023 Oct 27;51(19):10147-10161. doi: 10.1093/nar/gkad736. Nucleic Acids Res. 2023. PMID: 37738140 Free PMC article.
References
-
- Doudna JA; Charpentier E Genome Editing. The New Frontier of Genome Engineering with CRISPR-Cas9. Science 2014, 346, 1258096. - PubMed
-
- Stadtmauer EA; Fraietta JA; Davis MM; Cohen AD; Weber KL; Lancaster E; Mangan PA; Kulikovskaya I; Gupta M; Chen F; Tian L; Gonzalez VE; Xu J; Jung I; Melenhorst JJ; Plesa G; Shea J; Matlawski T; Cervini A; Gaymon AL; Desjardins S; Lamontagne A; Salas-Mckee J; Fesnak A; Siegel DL; Levine BL; Jadlowsky JK; Young RM; Chew A; Hwang W; Hexner EO; Carreno BM; Nobles CL; Bushman FD; Parker KR; Qi Y; Satpathy AT; Chang HY; Zhao Y; Lacey SF; June CH CRISPR-Engineered T Cells in Patients with Refractory Cancer. Science 2020, 367, eaba7365. - PMC - PubMed
