SalivaDirect: A simplified and flexible platform to enhance SARS-CoV-2 testing capacity
- PMID: 33521748
- PMCID: PMC7836249
- DOI: 10.1016/j.medj.2020.12.010
SalivaDirect: A simplified and flexible platform to enhance SARS-CoV-2 testing capacity
Abstract
Background: Scaling SARS-CoV-2 testing to meet demands of safe reopenings continues to be plagued by assay costs and supply chain shortages. In response, we developed SalivaDirect, which received Emergency Use Authorization (EUA) from the U.S. Food and Drug Administration (FDA).
Methods: We simplified our saliva-based diagnostic test by (1) not requiring collection tubes with preservatives, (2) replacing nucleic acid extraction with a simple enzymatic and heating step, and (3) testing specimens with a dualplex qRT-PCR assay. Moreover, we validated SalivaDirect with reagents and instruments from multiple vendors to minimize supply chain issues.
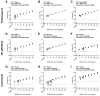
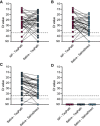
Findings: From our hospital cohort, we show a high positive agreement (94%) between saliva tested with SalivaDirect and nasopharyngeal swabs tested with a commercial qRT-PCR kit. In partnership with the National Basketball Association (NBA) and National Basketball Players Association (NBPA), we tested 3,779 saliva specimens from healthy individuals and detected low rates of invalid (0.3%) and false-positive (<0.05%) results.
Conclusions: We demonstrate that saliva is a valid alternative to swabs for SARS-CoV-2 screening and that SalivaDirect can make large-scale testing more accessible and affordable. Uniquely, we can designate other laboratories to use our sensitive, flexible, and simplified platform under our EUA (https://publichealth.yale.edu/salivadirect/).
Funding: This study was funded by the NBA and NBPA (N.D.G.), the Huffman Family Donor Advised Fund (N.D.G.), a Fast Grant from Emergent Ventures at the Mercatus Center at George Mason University (N.D.G.), the Yale Institute for Global Health (N.D.G.), and the Beatrice Kleinberg Neuwirth Fund (A.I.K.). C.B.F.V. is supported by NWO Rubicon 019.181EN.004.
Keywords: COVID-19; SARS-CoV-2; molecular testing; population screening; saliva.
© 2020 Elsevier Inc.
Conflict of interest statement
A.L.W. has received research funding through grants from Pfizer to Yale and has received consulting fees for participation in advisory boards for Pfizer. N.D.G. and A.L.W. have received research funding from Tempus to Yale to develop future versions of SalivaDirect. The remaining authors declare no competing interests.
Figures



References
-
- World Health Organization Coronavirus Disease (COVID-19) Situation Reports. World Health Organization. 2020. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situatio...
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

