Clusters of a Few Bound Cofilins Sever Actin Filaments
- PMID: 33524412
- PMCID: PMC8689643
- DOI: 10.1016/j.jmb.2021.166833
Clusters of a Few Bound Cofilins Sever Actin Filaments
Abstract
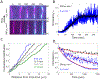
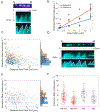
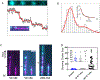
Cofilin is an essential actin filament severing protein that accelerates the assembly dynamics and turnover of actin networks by increasing the number of filament ends where subunits add and dissociate. It binds filament subunits stoichiometrically and cooperatively, forming clusters of contiguously-bound cofilin at sub-saturating occupancies. Filaments partially occupied with cofilin sever at boundaries between bare and cofilin-decorated segments. Imaging studies concluded that bound clusters must reach a critical size (Cc) of 13-100 cofilins to sever filaments. In contrast, structural and modeling studies suggest that a few or even a single cofilin can sever filaments, possibly with different severing rate constants. How clusters grow through the cooperative incorporation of additional cofilin molecules, specifically if they elongate asymmetrically or uniformly from both ends and if they are modulated by filament shape and external force, also lacks consensus. Here, using hydrodynamic flow to visualize individual actin filaments with TIRF microscopy, we found that neither flow-induced filament bending, tension, nor surface attachment conditions substantially affected the kinetics of cofilin binding to actin filaments. Clusters of bound cofilin preferentially extended toward filament pointed ends and displayed severing competency at small sizes (Cc < 3), with no detectable severing dependence on cluster size. These data support models in which small clusters of cofilin introduce local, but asymmetric, structural changes in actin filaments that promote filament severing with a rate constant that depends weakly on the size of the cluster.
Keywords: curvature; fluorescence; kinetics; microfluidics; tension.
Copyright © 2021. Published by Elsevier Ltd.
Conflict of interest statement
Declaration of Competing Interest The authors declare no competing interest.
Figures
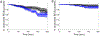
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

