Cellular response to spinal cord injury in regenerative and non-regenerative stages in Xenopus laevis
- PMID: 33526076
- PMCID: PMC7852093
- DOI: 10.1186/s13064-021-00152-2
Cellular response to spinal cord injury in regenerative and non-regenerative stages in Xenopus laevis
Abstract
Background: The efficient regenerative abilities at larvae stages followed by a non-regenerative response after metamorphosis in froglets makes Xenopus an ideal model organism to understand the cellular responses leading to spinal cord regeneration.
Methods: We compared the cellular response to spinal cord injury between the regenerative and non-regenerative stages of Xenopus laevis. For this analysis, we used electron microscopy, immunofluorescence and histological staining of the extracellular matrix. We generated two transgenic lines: i) the reporter line with the zebrafish GFAP regulatory regions driving the expression of EGFP, and ii) a cell specific inducible ablation line with the same GFAP regulatory regions. In addition, we used FACS to isolate EGFP+ cells for RNAseq analysis.
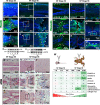
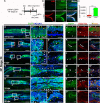
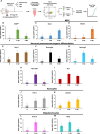
Results: In regenerative stage animals, spinal cord regeneration triggers a rapid sealing of the injured stumps, followed by proliferation of cells lining the central canal, and formation of rosette-like structures in the ablation gap. In addition, the central canal is filled by cells with similar morphology to the cells lining the central canal, neurons, axons, and even synaptic structures. Regeneration is almost completed after 20 days post injury. In non-regenerative stage animals, mostly damaged tissue was observed, without clear closure of the stumps. The ablation gap was filled with fibroblast-like cells, and deposition of extracellular matrix components. No reconstruction of the spinal cord was observed even after 40 days post injury. Cellular markers analysis confirmed these histological differences, a transient increase of vimentin, fibronectin and collagen was detected in regenerative stages, contrary to a sustained accumulation of most of these markers, including chondroitin sulfate proteoglycans in the NR-stage. The zebrafish GFAP transgenic line was validated, and we have demonstrated that is a very reliable and new tool to study the role of neural stem progenitor cells (NSPCs). RNASeq of GFAP::EGFP cells has allowed us to clearly demonstrate that indeed these cells are NSPCs. On the contrary, the GFAP::EGFP transgene is mainly expressed in astrocytes in non-regenerative stages. During regenerative stages, spinal cord injury activates proliferation of NSPCs, and we found that are mainly differentiated into neurons and glial cells. Specific ablation of these cells abolished proper regeneration, confirming that NSPCs cells are necessary for functional regeneration of the spinal cord.
Conclusions: The cellular response to spinal cord injury in regenerative and non-regenerative stages is profoundly different between both stages. A key hallmark of the regenerative response is the activation of NSPCs, which massively proliferate, and are differentiated into neurons to reconstruct the spinal cord. Also very notably, no glial scar formation is observed in regenerative stages, but a transient, glial scar-like structure is formed in non-regenerative stage animals.
Keywords: Gfap; Glial scar; NSPCs; Neurogenesis; Regeneration; Spinal cord; Xenopus; sox2.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures




Similar articles
-
Regeneration of Xenopus laevis spinal cord requires Sox2/3 expressing cells.Dev Biol. 2015 Dec 15;408(2):229-43. doi: 10.1016/j.ydbio.2015.03.009. Epub 2015 Mar 19. Dev Biol. 2015. PMID: 25797152 Free PMC article.
-
Cell Transplantation as a Method to Investigate Spinal Cord Regeneration in Regenerative and Nonregenerative Xenopus Stages.Cold Spring Harb Protoc. 2018 Dec 3;2018(12). doi: 10.1101/pdb.prot101006. Cold Spring Harb Protoc. 2018. PMID: 29769390
-
Genome-wide expression profile of the response to spinal cord injury in Xenopus laevis reveals extensive differences between regenerative and non-regenerative stages.Neural Dev. 2014 May 22;9:12. doi: 10.1186/1749-8104-9-12. Neural Dev. 2014. PMID: 24885550 Free PMC article.
-
The African clawed frog Xenopus laevis: A model organism to study regeneration of the central nervous system.Neurosci Lett. 2017 Jun 23;652:82-93. doi: 10.1016/j.neulet.2016.09.054. Epub 2016 Sep 29. Neurosci Lett. 2017. PMID: 27693567 Review.
-
Xenopus laevis as a Model Organism for the Study of Spinal Cord Formation, Development, Function and Regeneration.Front Neural Circuits. 2017 Nov 23;11:90. doi: 10.3389/fncir.2017.00090. eCollection 2017. Front Neural Circuits. 2017. PMID: 29218002 Free PMC article. Review.
Cited by
-
Neurotrauma-From Injury to Repair: Clinical Perspectives, Cellular Mechanisms and Promoting Regeneration of the Injured Brain and Spinal Cord.Biomedicines. 2024 Mar 13;12(3):643. doi: 10.3390/biomedicines12030643. Biomedicines. 2024. PMID: 38540256 Free PMC article. Review.
-
Xenopus laevis neural stem progenitor cells exhibit a transient metabolic shift toward glycolysis during spinal cord regeneration.Front Cell Dev Biol. 2025 Jan 29;13:1529093. doi: 10.3389/fcell.2025.1529093. eCollection 2025. Front Cell Dev Biol. 2025. PMID: 39944083 Free PMC article.
-
Animal Models of Spinal Cord Injury.Biomedicines. 2025 Jun 10;13(6):1427. doi: 10.3390/biomedicines13061427. Biomedicines. 2025. PMID: 40564146 Free PMC article. Review.
-
The Stress Response of the Holothurian Central Nervous System: A Transcriptomic Analysis.Int J Mol Sci. 2022 Nov 2;23(21):13393. doi: 10.3390/ijms232113393. Int J Mol Sci. 2022. PMID: 36362181 Free PMC article.
-
Mitochondrial function in spinal cord injury and regeneration.Cell Mol Life Sci. 2022 Apr 13;79(5):239. doi: 10.1007/s00018-022-04261-x. Cell Mol Life Sci. 2022. PMID: 35416520 Free PMC article. Review.
References
-
- Diaz Quiroz JF, Echeverri K. Spinal cord regeneration: where fish, frogs and salamanders lead the way, can we follow? Biochem J. 2013;451(3):353–364. - PubMed
-
- Cardozo MJ, Mysiak K, Becker T, Becker CG. Reduce, reuse, recycle - developmental signals in spinal cord regeneration. Dev Biol. 2017;432(1):53–62. - PubMed
-
- Marichal N, Reali C, Rehermann MI, Trujillo-Cenóz O, Russo RE. Progenitors in the Ependyma of the spinal cord: a potential resource for self-repair after injury. Adv Exp Med Biol. 2017;1015:241–264. - PubMed
-
- Lee-Liu D, Méndez-Olivos EE, Muñoz R, Larraín J. The African clawed frog Xenopus laevis: a model organism to study regeneration of the central nervous system. Neurosci Lett. 2017;652:82–93. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

