Genomic Insights into the Origin and Evolution of Molluscan Red-Bloodedness in the Blood Clam Tegillarca granosa
- PMID: 33528571
- PMCID: PMC8136487
- DOI: 10.1093/molbev/msab030
Genomic Insights into the Origin and Evolution of Molluscan Red-Bloodedness in the Blood Clam Tegillarca granosa
Erratum in
-
Erratum to: Genomic Insights into the Origin and Evolution of Molluscan Red-Bloodedness in the Blood Clam Tegillarca granosa.Mol Biol Evol. 2021 Jul 29;38(8):3494. doi: 10.1093/molbev/msab126. Mol Biol Evol. 2021. PMID: 34046673 Free PMC article. No abstract available.
Abstract
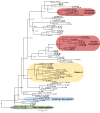
Blood clams differ from their molluscan kins by exhibiting a unique red-blood (RB) phenotype; however, the genetic basis and biochemical machinery subserving this evolutionary innovation remain unclear. As a fundamental step toward resolving this mystery, we presented the first chromosome-level genome and comprehensive transcriptomes of the blood clam Tegillarca granosa for an integrated genomic, evolutionary, and functional analyses of clam RB phenotype. We identified blood clam-specific and expanded gene families, as well as gene pathways that are of RB relevant. Clam-specific RB-related hemoglobins (Hbs) showed close phylogenetic relationships with myoglobins (Mbs) of blood clam and other molluscs without the RB phenotype, indicating that clam-specific Hbs were likely evolutionarily derived from the Mb lineage. Strikingly, similar to vertebrate Hbs, blood clam Hbs were present in a form of gene cluster. Despite the convergent evolution of Hb clusters in blood clam and vertebrates, their Hb clusters may have originated from a single ancestral Mb-like gene as evidenced by gene phylogeny and synteny analysis. A full suite of enzyme-encoding genes for heme synthesis was identified in blood clam, with prominent expression in hemolymph and resembling those in vertebrates, suggesting a convergence of both RB-related Hb and heme functions in vertebrates and blood clam. RNA interference experiments confirmed the functional roles of Hbs and key enzyme of heme synthesis in the maintenance of clam RB phenotype. The high-quality genome assembly and comprehensive transcriptomes presented herein serve new genomic resources for the super-diverse phylum Mollusca, and provide deep insights into the origin and evolution of invertebrate RB.
Keywords: blood clam; genome sequencing; heme biosynthesis; hemoglobin evolution; red-bloodedness.
© The Author(s) 2021. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures





References
-
- Afiati N. 1999. Cytoplasmic granules in the red blood cells and the karyotype of rounded ecomorph of Anadara granosa (L.)(Bivalvia: arcidae) from Central Java. Indonesia. Majalah Ilmu Kelautan IV. 14:51–59.
-
- Ajioka RS, Phillips JD, Kushner JP.. 2006. Biosynthesis of heme in mammals. Biochim Biophys Acta 1763(7):723–736. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

