Increased IGFBP7 Expression Correlates with Poor Prognosis and Immune Infiltration in Gastric Cancer
- PMID: 33531979
- PMCID: PMC7847654
- DOI: 10.7150/jca.50370
Increased IGFBP7 Expression Correlates with Poor Prognosis and Immune Infiltration in Gastric Cancer
Abstract
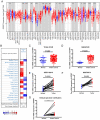
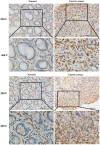
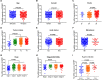
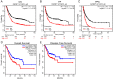
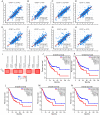
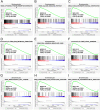
Background: Insulin-like growth factor binding protein-7 (IGFBP7) contributes to multiple biological processes in various tumors. However, the role of IGFBP7 in gastric cancer (GC) is still undetermined. The study aims to explore the role of IGFBP7 in GC via an integrated bioinformatics analysis. Methods: IGFBP7 expression levels in GC and its normal gastric tissues were analyzed using multiple databases, including the Tumor Immune Estimation Resource (TIMER), Oncomine, The Cancer Genome Atlas (TCGA) and Gene Expression Omnibus (GEO) databases, as well as by our clinical gastric specimens. The methylation analysis was conducted with MEXPRESS, UALCAN and Xena online tools. The survival analysis was conducted using the Kaplan-Meier Plotter and Gene Expression Profiling Interactive Analysis (GEPIA) databases. Coexpressed genes of IGFBP7 were selected with the cBioPortal tool and enrichment analysis was conducted with the clusterProfiler package in R software. Gene set enrichment analysis (GSEA) was performed to explore the IGFBP7-related biological processes involved in GC. Correlations between IGFBP7 and immune cell infiltrates were analyzed using the TIMER database. Results: IGFBP7 expression was significantly upregulated in GC and correlated with stage, grade, tumor status and Helicobacter pylori infection. High IGFBP7 expression and low IGFBP7 methylation levels were significantly associated with short survival of patients with GC. Univariate and multivariate analyses revealed that IGFBP7 was an independent risk factor for GC. The coexpressed genes LHFPL6, SEPTIN4, HSPB2, LAYN and GGT5 predicted unfavorable outcomes of GC. Enrichment analysis showed that the coexpressed genes were involved in extracellular matrix (ECM)-related processes. GSEA indicated that IGFBP7 was positively related to ECM and inflammation-related pathways. TIMER analysis indicated that the mRNA level of IGFBP7 was strongly correlated with genes related to various infiltrating immune cells in GC, especially with gene markers of tumor associated macrophages (TAMs). Conclusions: Increased IGFBP7 expression correlates with poor prognosis and immune cell infiltration in GC, which might be a potential biomarker for the diagnosis of GC.
Keywords: IGFBP7; bioinformatics analysis; gastric cancer; immune infiltration; prognosis.
© The author(s).
Conflict of interest statement
Competing Interests: The authors have declared that no competing interest exists.
Figures

References
-
- Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. - PubMed
-
- Choi IJ, Kim CG, Lee JY, Kim YI, Kook MC, Park B, Joo J. Family History of Gastric Cancer and Helicobacter pylori Treatment. N Engl J Med. 2020;382(5):427–436. - PubMed
-
- Fukase K, Kato M, Kikuchi S, Inoue K, Uemura N, Okamoto S, Terao S, Amagai K, Hayashi S, Asaka M. Effect of eradication of Helicobacter pylori on incidence of metachronous gastric carcinoma after endoscopic resection of early gastric cancer: an open-label, randomised controlled trial. Lancet. 2008;372(9636):392–397. - PubMed
-
- Oh Y. IGFBPs and neoplastic models. New concepts for roles of IGFBPs in regulation of cancer cell growth. Endocrine. 1997;7(1):111–113. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

