Multiomics Analysis Provides Insight into the Laboratory Evolution of Escherichia coli toward the Metabolic Usage of Fluorinated Indoles
- PMID: 33532571
- PMCID: PMC7844855
- DOI: 10.1021/acscentsci.0c00679
Multiomics Analysis Provides Insight into the Laboratory Evolution of Escherichia coli toward the Metabolic Usage of Fluorinated Indoles
Abstract
Organofluorine compounds are known to be toxic to a broad variety of living beings in different habitats, and chemical fluorination has been historically exploited by mankind for the development of therapeutic drugs or agricultural pesticides. On the other hand, several studies so far have demonstrated that, under appropriate conditions, living systems (in particular bacteria) can tolerate the presence of fluorinated molecules (e.g., amino acids analogues) within their metabolism and even repurpose them as alternative building blocks for the synthesis of cellular macromolecules such as proteins. Understanding the molecular mechanism behind these phenomena would greatly advance approaches to the biotechnological synthesis of recombinant proteins and peptide drugs. However, information about the metabolic effects of long-term exposure of living cells to fluorinated amino acids remains scarce. Hereby, we report the long-term propagation of Escherichia coli (E. coli) in an artificially fluorinated habitat that yielded two strains naturally adapted to live on fluorinated amino acids. In particular, we applied selective pressure to force a tryptophan (Trp)-auxotrophic strain to use either 4- or 5-fluoroindole as essential precursors for the in situ synthesis of Trp analogues, followed by their incorporation in the cellular proteome. We found that full adaptation to both fluorinated Trp analogues requires a low number of genetic mutations but is accompanied by large rearrangements in regulatory networks, membrane integrity, and quality control of protein folding. These findings highlight the cellular mechanisms behind the adaptation to unnatural amino acids and provide the molecular foundation for bioengineering of novel microbial strains for synthetic biology and biotechnology.
© 2020 American Chemical Society.
Conflict of interest statement
The authors declare no competing financial interest.
Figures

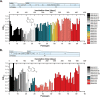



References
-
- Harper D. B.; O’Hagan D.; Murphy C. D. Fluorinated Natural Products: Occurrence and Biosynthesis. Handbook of Environmental Chemistry 2003, 3, 141–169. 10.1007/b10454. - DOI
Grants and funding
LinkOut - more resources
Full Text Sources

