A scaffold lncRNA shapes the mitosis to meiosis switch
- PMID: 33536434
- PMCID: PMC7859202
- DOI: 10.1038/s41467-021-21032-7
A scaffold lncRNA shapes the mitosis to meiosis switch
Abstract
Long non-coding RNAs (lncRNAs) contribute to the regulation of gene expression in response to intra- or extracellular signals but the underlying molecular mechanisms remain largely unexplored. Here, we identify an uncharacterized lncRNA as a central player in shaping the meiotic gene expression program in fission yeast. We report that this regulatory RNA, termed mamRNA, scaffolds the antagonistic RNA-binding proteins Mmi1 and Mei2 to ensure their reciprocal inhibition and fine tune meiotic mRNA degradation during mitotic growth. Mechanistically, mamRNA allows Mmi1 to target Mei2 for ubiquitin-mediated downregulation, and conversely enables accumulating Mei2 to impede Mmi1 activity, thereby reinforcing the mitosis to meiosis switch. These regulations also occur within a unique Mmi1-containing nuclear body, positioning mamRNA as a spatially-confined sensor of Mei2 levels. Our results thus provide a mechanistic basis for the mutual control of gametogenesis effectors and further expand our vision of the regulatory potential of lncRNAs.
Conflict of interest statement
The authors declare no competing interests.
Figures



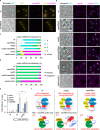
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

