Sea Urchin as a Universal Model for Studies of Gene Networks
- PMID: 33552139
- PMCID: PMC7854572
- DOI: 10.3389/fgene.2020.627259
Sea Urchin as a Universal Model for Studies of Gene Networks
Abstract
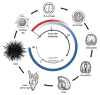
The purple sea urchin Strongylocentrotus purpuratus has been used for over 150 years as a model organism in developmental biology. Using this model species, scientists have been able to describe, in detail, the mechanisms of cell cycle control and cell adhesion, fertilization, calcium signaling, cell differentiation, and death. Massive parallel sequencing of the sea urchin genome enabled the deciphering of the main components of gene regulatory networks during the activation of embryonic signaling pathways. This knowledge helped to extrapolate aberrations in somatic cells that may lead to diseases, including cancer in humans. Furthermore, since many, if not all, developmental signaling pathways were shown to be controlled by non-coding RNAs (ncRNAs), the sea urchin organism represents an attractive experimental model. In this review, we discuss the main discoveries in the genetics, genomics, and transcriptomics of sea urchins during embryogenesis with the main focus on the role of ncRNAs. This information may be useful for comparative studies between different organisms, and may help identify new regulatory networks controlled by ncRNAs.
Keywords: cell signaling; gene expression; genomics; long non-coding RNA; sea urchin.
Copyright © 2021 Adonin, Drozdov and Barlev.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Ocean acidification research in the 'post-genomic' era: Roadmaps from the purple sea urchin Strongylocentrotus purpuratus.Comp Biochem Physiol A Mol Integr Physiol. 2015 Jul;185:33-42. doi: 10.1016/j.cbpa.2015.03.007. Epub 2015 Mar 13. Comp Biochem Physiol A Mol Integr Physiol. 2015. PMID: 25773301 Review.
-
RNA deep sequencing reveals differential microRNA expression during development of sea urchin and sea star.PLoS One. 2011;6(12):e29217. doi: 10.1371/journal.pone.0029217. Epub 2011 Dec 28. PLoS One. 2011. PMID: 22216218 Free PMC article.
-
Methods for karyotyping and for localization of developmentally relevant genes on the chromosomes of the purple sea urchin, Strongylocentrotus purpuratus.Biol Bull. 2009 Dec;217(3):306-12. doi: 10.1086/BBLv217n3p306. Biol Bull. 2009. PMID: 20040754
-
Global patterns of enhancer activity during sea urchin embryogenesis assessed by eRNA profiling.Genome Res. 2021 Sep;31(9):1680-1692. doi: 10.1101/gr.275684.121. Epub 2021 Jul 30. Genome Res. 2021. PMID: 34330790 Free PMC article.
-
The painted sea urchin, Lytechinus pictus, as a genetically-enabled developmental model.Methods Cell Biol. 2019;150:105-123. doi: 10.1016/bs.mcb.2018.11.010. Epub 2019 Feb 10. Methods Cell Biol. 2019. PMID: 30777173 Free PMC article. Review.
Cited by
-
Pattern of Repetitive Element Transcription Segregate Cell Lineages during the Embryogenesis of Sea Urchin Strongylocentrotus purpuratus.Biomedicines. 2021 Nov 21;9(11):1736. doi: 10.3390/biomedicines9111736. Biomedicines. 2021. PMID: 34829966 Free PMC article.
-
Exploring the Bioactive Potential of Pisolithus (Basidiomycota): Comprehensive Insights into Antimicrobial, Anticancer, and Antioxidant Properties for Innovative Applications.Microorganisms. 2024 Feb 23;12(3):450. doi: 10.3390/microorganisms12030450. Microorganisms. 2024. PMID: 38543501 Free PMC article. Review.
-
PFAS Compounds PFOA and Gen X are Teratogenic to Sea Urchin Embryos.bioRxiv [Preprint]. 2024 Nov 21:2024.11.21.624751. doi: 10.1101/2024.11.21.624751. bioRxiv. 2024. Update in: Dev Biol. 2025 Sep;525:139-154. doi: 10.1016/j.ydbio.2025.06.004. PMID: 39605628 Free PMC article. Updated. Preprint.
-
Taxonomic Diversity, Predicted Metabolic Pathway, and Interaction Pattern of Bacterial Community in Sea Urchin Anthocidaris crassispina.Microorganisms. 2024 Oct 20;12(10):2094. doi: 10.3390/microorganisms12102094. Microorganisms. 2024. PMID: 39458402 Free PMC article.
-
Similarities in the induction of the intracellular pathogen response in Caenorhabditis elegans and the type I interferon response in mammals.Bioessays. 2023 Nov;45(11):e2300097. doi: 10.1002/bies.202300097. Epub 2023 Sep 4. Bioessays. 2023. PMID: 37667453 Free PMC article.
References
-
- Aristotle (1942). Generation of animals. Peck A. L. Loeb Classical Library 366 Cambridge, MA: Harvard University Press.
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

