Identifying Wild Versus Cultivated Gene-Alleles Conferring Seed Coat Color and Days to Flowering in Soybean
- PMID: 33557103
- PMCID: PMC7913812
- DOI: 10.3390/ijms22041559
Identifying Wild Versus Cultivated Gene-Alleles Conferring Seed Coat Color and Days to Flowering in Soybean
Abstract
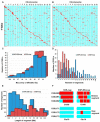
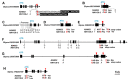
Annual wild soybean (G. soja) is the ancestor of the cultivated soybean (G. max). To reveal the genetic changes from soja to max, an improved wild soybean chromosome segment substitution line (CSSL) population, SojaCSSLP5, composed of 177 CSSLs with 182 SSR markers (SSR-map), was developed based on SojaCSSLP1 generated from NN1138-2(max)×N24852(soja). The SojaCSSLP5 was genotyped further through whole-genome resequencing, resulting in a physical map with 1366 SNPLDBs (SNP linkage-disequilibrium blocks), which are composed of more markers/segments, shorter marker length and more recombination breakpoints than the SSR-map and caused 721 new wild substituted segments. Using the SNPLDB-map, two loci co-segregating with seed-coat color (SCC) and six loci for days to flowering (DTF) with 88.02% phenotypic contribution were identified. Integrated with parental RNA-seq and DNA-resequencing, two SCC and six DTF candidate genes, including three previously cloned (G, E2 and GmPRR3B) and five newly detected ones, were predicted and verified at nucleotide mutant level, and then demonstrated with the consistency between gene-alleles and their phenotypes in SojaCSSLP5. In total, six of the eight genes were identified with the parental allele-pairs coincided to those in 303 germplasm accessions, then were further demonstrated by the consistency between gene-alleles and germplasm phenotypes. Accordingly, the CSSL population integrated with parental DNA and RNA sequencing data was demonstrated to be an efficient platform in identifying candidate wild vs. cultivated gene-alleles.
Keywords: SNP linkage disequilibrium block (SNPLDB); annual wild soybean (G. soja Sieb. and Zucc.); chromosome segment substitution line (CSSL); cultivated soybean (G. max (L.) Merr.); days to flowering; seed coat color; whole-genome re-sequencing.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Detecting the QTL-allele system conferring flowering date in a nested association mapping population of soybean using a novel procedure.Theor Appl Genet. 2017 Nov;130(11):2297-2314. doi: 10.1007/s00122-017-2960-y. Epub 2017 Aug 10. Theor Appl Genet. 2017. PMID: 28799029
-
Identification of novel loci associated with maturity and yield traits in early maturity soybean plant introduction lines.BMC Genomics. 2018 Mar 1;19(1):167. doi: 10.1186/s12864-018-4558-4. BMC Genomics. 2018. PMID: 29490606 Free PMC article.
-
Dissecting genomic hotspots underlying seed protein, oil, and sucrose content in an interspecific mapping population of soybean using high-density linkage mapping.Plant Biotechnol J. 2018 Nov;16(11):1939-1953. doi: 10.1111/pbi.12929. Epub 2018 May 16. Plant Biotechnol J. 2018. PMID: 29618164 Free PMC article.
-
A platform for soybean molecular breeding: the utilization of core collections for food security.Plant Mol Biol. 2013 Sep;83(1-2):41-50. doi: 10.1007/s11103-013-0076-6. Epub 2013 May 25. Plant Mol Biol. 2013. PMID: 23708950 Free PMC article. Review.
-
Divergence of flowering genes in soybean.J Biosci. 2012 Nov;37(5):857-70. doi: 10.1007/s12038-012-9252-0. J Biosci. 2012. PMID: 23107921 Review.
Cited by
-
The Key to the Future Lies in the Past: Insights from Grain Legume Domestication and Improvement Should Inform Future Breeding Strategies.Plant Cell Physiol. 2022 Nov 22;63(11):1554-1572. doi: 10.1093/pcp/pcac086. Plant Cell Physiol. 2022. PMID: 35713290 Free PMC article. Review.
-
Dissection of the E8 locus in two early maturing Canadian soybean populations.Front Plant Sci. 2024 Feb 8;15:1329065. doi: 10.3389/fpls.2024.1329065. eCollection 2024. Front Plant Sci. 2024. PMID: 38390301 Free PMC article.
-
Identification of genetic loci conferring seed coat color based on a high-density map in soybean.Front Plant Sci. 2022 Aug 1;13:968618. doi: 10.3389/fpls.2022.968618. eCollection 2022. Front Plant Sci. 2022. PMID: 35979081 Free PMC article.
-
Research advances on the hard seededness trait of soybean and the underlying regulatory mechanisms.Front Plant Sci. 2024 Jun 26;15:1419962. doi: 10.3389/fpls.2024.1419962. eCollection 2024. Front Plant Sci. 2024. PMID: 38988633 Free PMC article. Review.
-
Identification of candidate genes for soybean seed coat-related traits using QTL mapping and GWAS.Front Plant Sci. 2023 Jun 13;14:1190503. doi: 10.3389/fpls.2023.1190503. eCollection 2023. Front Plant Sci. 2023. PMID: 37384360 Free PMC article.
References
-
- Broich S.L., Palmer R.G. A cluster analysis of wild and domesticated soybean phenotypes. Euphytica. 1980;29:23–32. doi: 10.1007/BF00037246. - DOI
-
- Prince S.J., Song L., Qiu D., Maldonado Dos Santos J.V., Chai C., Joshi T., Patil G., Valliyodan B., Vuong T.D., Murphy M., et al. Genetic variants in root architecture-related genes in a Glycine soja accession, a potential resource to improve cultivated soybean. BMC Genom. 2015;16:132. doi: 10.1186/s12864-015-1334-6. - DOI - PMC - PubMed
-
- La T., Large E., Taliercio E., Song Q., Gillman J.D., Xu D., Nguyen H.T., Shannon G., Scaboo A. Characterization of Select Wild Soybean Accessions in the USDA Germplasm Collection for Seed Composition and Agronomic Traits. Crop. Sci. 2019;58:1–19. doi: 10.2135/cropsci2017.08.0514. - DOI
-
- Wang W., He Q., Yang H., Xiang S., Zhao T., Gai J. Development of a chromosome segment substitution line population with wild soybean (Glycine soja Sieb. et Zucc.) as donor parent. Euphytica. 2013;189:293–307. doi: 10.1007/s10681-012-0817-7. - DOI
-
- Yang H.Y., Wang W.B., He Q.Y., Xiang S.H., Tian D., Zhao T.J., Gai J.Y. Chromosome segment detection for seed size and shape traits using an improved population of wild soybean chromosome segment substitution lines. Physiol. Mol. Biol. Plants. 2017;23:877–889. doi: 10.1007/s12298-017-0468-1. - DOI - PMC - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

