SPRTN protease-cleaved MRE11 decreases DNA repair and radiosensitises cancer cells
- PMID: 33558481
- PMCID: PMC7870818
- DOI: 10.1038/s41419-021-03437-w
SPRTN protease-cleaved MRE11 decreases DNA repair and radiosensitises cancer cells
Abstract
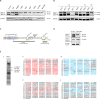
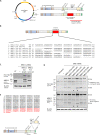
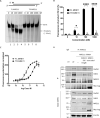
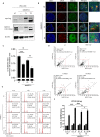
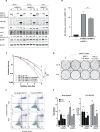
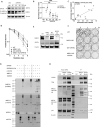
The human MRE11/RAD50/NBS1 (MRN) complex plays a crucial role in sensing and repairing DNA DSB. MRE11 possesses dual 3'-5' exonuclease and endonuclease activity and forms the core of the multifunctional MRN complex. We previously identified a C-terminally truncated form of MRE11 (TR-MRE11) associated with post-translational MRE11 degradation. Here we identified SPRTN as the essential protease for the formation of TR-MRE11 and characterised the role of this MRE11 form in its DNA damage response (DDR). Using tandem mass spectrometry and site-directed mutagenesis, the SPRTN-dependent cleavage site for MRE11 was identified between 559 and 580 amino acids. Despite the intact interaction of TR-MRE11 with its constitutive core complex proteins RAD50 and NBS1, both nuclease activities of truncated MRE11 were dramatically reduced due to its deficient binding to DNA. Furthermore, lack of the MRE11 C-terminal decreased HR repair efficiency, very likely due to abolished recruitment of TR-MRE11 to the sites of DNA damage, which consequently led to increased cellular radiosensitivity. The presence of this DNA repair-defective TR-MRE11 could explain our previous finding that the high MRE11 protein expression by immunohistochemistry correlates with improved survival following radical radiotherapy in bladder cancer patients.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

