Loss of KDM4B exacerbates bone-fat imbalance and mesenchymal stromal cell exhaustion in skeletal aging
- PMID: 33571444
- PMCID: PMC8178178
- DOI: 10.1016/j.stem.2021.01.010
Loss of KDM4B exacerbates bone-fat imbalance and mesenchymal stromal cell exhaustion in skeletal aging
Abstract
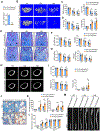
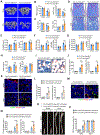
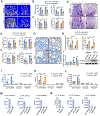
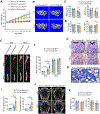
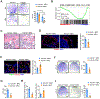
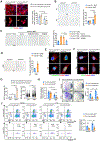
Skeletal aging is a complex process, characterized by a decrease in bone formation, an increase in marrow fat, and stem cell exhaustion. Loss of H3K9me3, a heterochromatin mark, has been proposed to be associated with aging. Here, we report that loss of KDM4B in mesenchymal stromal cells (MSCs) exacerbated skeletal aging and osteoporosis by reducing bone formation and increasing marrow adiposity via increasing H3K9me3. KDM4B epigenetically coordinated β-catenin/Smad1-mediated transcription by removing repressive H3K9me3. Importantly, KDM4B ablation impaired MSC self-renewal and promoted MSC exhaustion by inducing senescence-associated heterochromatin foci formation, providing a mechanistic explanation for stem cell exhaustion with aging. Moreover, while KDM4B was required for parathyroid hormone-mediated bone anabolism, KDM4B depletion accelerated bone loss and marrow adiposity induced by a high-fat diet. Our results suggest that the epigenetic rejuvenation and reversing bone-fat imbalance might be new strategies for preventing and treating skeletal aging and osteoporosis by activating KDM4B in MSCs.
Keywords: bone marrow adiposity; bone metabolism; mesenchymal stem cells; mesenchymal stromal cells; osteoporosis; parathyroid hormone; senescence-associated heterochromatin foci; skeletal aging; stem cell self-renewal.
Copyright © 2021 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures

References
-
- Burge R, Dawson-Hughes B, Solomon DH, Wong JB, King A, and Tosteson A (2007). Incidence and Economic Burden of Osteoporosis-Related Fractures in the United States, 2005–2025. J Bone Miner Res 22, 465–475. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

