A Chromosome-Level Genome Assembly of the Dark Sleeper Odontobutis potamophila
- PMID: 33576781
- PMCID: PMC7883661
- DOI: 10.1093/gbe/evaa271
A Chromosome-Level Genome Assembly of the Dark Sleeper Odontobutis potamophila
Abstract
The dark sleeper, Odontobutis potamophila, is a commercially valuable fish that widely distributed in China and Southeast Asia countries. The phenomenon of sexual dimorphism in growth is conspicuous, which the males grow substantially larger and faster than the females. However, the high-quality genome resources for gaining insight into sex-determining mechanisms to develop sex-control breeding are still lacking. Here, a chromosomal-level genome assembly of O. potamophila was generated from a combination of Illumina reads, 10× Genomics sequencing, and Hi-C chromatin interaction sequencing. The assembled genome was 1,134.62 Mb with a contig N50 of 22.25 Mb and a scaffold N50 of 24.85 Mb, representing 94.4% completeness (Benchmarking Universal Single-Copy Orthologs). Using Hi-C data, 96.49% of the total contig bases were anchored to the 22 chromosomes, with a contig N50 of 22.25 Mb and a scaffold N50 of 47.68 Mb. Approximately 54.18% of the genome were identified as repetitive elements, and 23,923 protein-coding genes were annotated in the genome. The assembled genome can be used as a valuable resource for molecular breeding and functional studies of O. potamophila in the future.
Keywords: Odontobutis potamophila; gene annotation; sex determination; whole-genome sequence.
© The Author(s) 2021. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
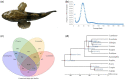
Figures
Similar articles
-
Genome sequence of the barred knifejaw Oplegnathus fasciatus (Temminck & Schlegel, 1844): the first chromosome-level draft genome in the family Oplegnathidae.Gigascience. 2019 Mar 1;8(3):giz013. doi: 10.1093/gigascience/giz013. Gigascience. 2019. PMID: 30715332 Free PMC article.
-
Chromosome-level genome assembly of the greenfin horse-faced filefish (Thamnaconus septentrionalis) using Oxford Nanopore PromethION sequencing and Hi-C technology.Mol Ecol Resour. 2020 Jul;20(4):1069-1079. doi: 10.1111/1755-0998.13183. Epub 2020 Jul 9. Mol Ecol Resour. 2020. PMID: 32390337
-
Chromosome-level genome assembly and annotation of Chinese herring (Ilisha elongata).Sci Data. 2025 Apr 21;12(1):668. doi: 10.1038/s41597-025-04790-7. Sci Data. 2025. PMID: 40258812 Free PMC article.
-
Chromosome-Level Genome Assembly and Annotation of a Sciaenid Fish, Argyrosomus japonicus.Genome Biol Evol. 2021 Feb 3;13(2):evaa246. doi: 10.1093/gbe/evaa246. Genome Biol Evol. 2021. PMID: 33484557 Free PMC article.
-
Chromosome-level reference genome of the Siamese fighting fish Betta splendens, a model species for the study of aggression.Gigascience. 2018 Nov 1;7(11):giy087. doi: 10.1093/gigascience/giy087. Gigascience. 2018. PMID: 30010754 Free PMC article.
Cited by
-
Chromosome-level genome assembly and annotation of largemouth bronze gudgeon (Coreius guichenoti).Sci Data. 2025 Jan 15;12(1):76. doi: 10.1038/s41597-025-04416-y. Sci Data. 2025. PMID: 39814798 Free PMC article.
-
Multi-omics Investigation of Freeze Tolerance in the Amur Sleeper, an Aquatic Ectothermic Vertebrate.Mol Biol Evol. 2023 Mar 4;40(3):msad040. doi: 10.1093/molbev/msad040. Mol Biol Evol. 2023. PMID: 36805964 Free PMC article.
-
Effects of Quercetin on the Intestinal Microflora of Freshwater Dark Sleeper Odontobutis potamophila.Antioxidants (Basel). 2022 Oct 12;11(10):2015. doi: 10.3390/antiox11102015. Antioxidants (Basel). 2022. PMID: 36290739 Free PMC article.
-
Effect of quercetin on muscle growth and antioxidant status of the dark sleeper Odontobutis potamophila.Front Genet. 2022 Jul 25;13:938526. doi: 10.3389/fgene.2022.938526. eCollection 2022. Front Genet. 2022. PMID: 35957695 Free PMC article.
-
Chromosome-level genome assembly of largemouth bass (Micropterus salmoides) using PacBio and Hi-C technologies.Sci Data. 2022 Aug 6;9(1):482. doi: 10.1038/s41597-022-01601-1. Sci Data. 2022. PMID: 35933561 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

