Host-Dependent Clustering of Campylobacter Strains From Small Mammals in Finland
- PMID: 33584588
- PMCID: PMC7873845
- DOI: 10.3389/fmicb.2020.621490
Host-Dependent Clustering of Campylobacter Strains From Small Mammals in Finland
Abstract
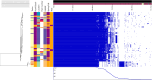
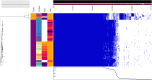
Small mammals are known to carry Campylobacter spp.; however, little is known about the genotypes and their role in human infections. We studied intestinal content from small wild mammals collected in their natural habitats in Finland in 2010-2017, and in close proximity to 40 pig or cattle farms in 2017. The animals were trapped using traditional Finnish metal snap traps. Campylobacter spp. were isolated from the intestinal content using direct plating on mCCDA. A total of 19% of the captured wild animals (n = 577) and 41% of the pooled farm samples (n = 227) were positive for C. jejuni, which was the only Campylobacter species identified. The highest prevalence occurred in yellow-necked mice (Apodemus flavicollis) and bank voles (Myodes glareolus) which carried Campylobacter spp. in 66.3 and 63.9% of the farm samples and 41.5 and 24.4% of individual animals trapped from natural habitats, respectively. Interestingly, all house mouse (Mus musculus) and shrew (Sorex spp.) samples were negative for Campylobacter spp. C. jejuni isolates (n = 145) were further characterized by whole-genome sequencing. Core genome multilocus sequence typing (cgMLST) clustering showed that mouse and vole strains were separated from the rest of the C. jejuni population (636 and 671 allelic differences, 94 and 99% of core loci, respectively). Very little or no alleles were shared with C. jejuni genomes described earlier from livestock or human isolates. FastANI results further indicated that C. jejuni strains from voles are likely to represent a new previously undescribed species or subspecies of Campylobacter. Core-genome phylogeny showed that there was no difference between isolates originating from the farm and wild captured animals. Instead, the phylogeny followed the host species-association. There was some evidence (one strain each) of livestock-associated C. jejuni occurring in a farm-caught A. flavicollis and a brown rat (Rattus norvegicus), indicating that although small mammals may not be the original reservoir of Campylobacter colonizing livestock, they may sporadically carry C. jejuni strains occurring mainly in livestock and be associated with disease in humans.
Keywords: Campylobacter jejuni; comparative genomics; mouse; phylogeny; rodent; shrew; vole.
Copyright © 2021 Olkkola, Rossi, Jaakkonen, Simola, Tikkanen, Hakkinen, Tuominen, Huitu, Niemimaa, Henttonen and Kivistö.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
Small mammals as carriers of zoonotic bacteria on pig and cattle farms - Prevalence and risk of exposure in an integrative approach.Prev Vet Med. 2024 Aug;229:106228. doi: 10.1016/j.prevetmed.2024.106228. Epub 2024 May 21. Prev Vet Med. 2024. PMID: 38850871
-
Prevalence and genotype diversity of Campylobacter jejuni in hunted reared pheasants (Phasianus colchicus) in Finland.Acta Vet Scand. 2023 Aug 1;65(1):36. doi: 10.1186/s13028-023-00698-7. Acta Vet Scand. 2023. PMID: 37528488 Free PMC article.
-
Campylobacter jejuni and Campylobacter coli in wild birds on Danish livestock farms.Acta Vet Scand. 2016 Feb 3;58:11. doi: 10.1186/s13028-016-0192-9. Acta Vet Scand. 2016. PMID: 26842400 Free PMC article.
-
Campylobacter species and genotype distribution in Finnish beef liver - Retail liver juice ideal for isolation and quantification.Int J Food Microbiol. 2024 Feb 2;411:110524. doi: 10.1016/j.ijfoodmicro.2023.110524. Epub 2023 Dec 14. Int J Food Microbiol. 2024. PMID: 38118359
-
Population Biology and Comparative Genomics of Campylobacter Species.Curr Top Microbiol Immunol. 2021;431:59-78. doi: 10.1007/978-3-030-65481-8_3. Curr Top Microbiol Immunol. 2021. PMID: 33620648 Review.
Cited by
-
Detection of multiple human enteropathogens in Norway rats (Rattus norvegicus) from an under-resourced neighborhood of Vancouver, British Columbia.PLoS Negl Trop Dis. 2023 Oct 16;17(10):e0011669. doi: 10.1371/journal.pntd.0011669. eCollection 2023 Oct. PLoS Negl Trop Dis. 2023. PMID: 37844114 Free PMC article.
-
Intestinal Colonization by Campylobacter jejuni, Clostridium difficile, and Clostridium perfringens among Commensal Rattus norvegicus in the Urban Areas of Tehran, Iran.Biomed Res Int. 2024 Mar 27;2024:2929315. doi: 10.1155/2024/2929315. eCollection 2024. Biomed Res Int. 2024. PMID: 38572169 Free PMC article.
-
Potentially zoonotic pathogens and parasites in opportunistically sourced urban brown rats (Rattus norvegicus) in and around Helsinki, Finland, 2018 to 2023.Euro Surveill. 2024 Oct;29(40):2400031. doi: 10.2807/1560-7917.ES.2024.29.40.2400031. Euro Surveill. 2024. PMID: 39364602 Free PMC article.
-
A Systematic Review on the Role of Wildlife as Carriers and Spreaders of Campylobacter spp.Animals (Basel). 2023 Apr 13;13(8):1334. doi: 10.3390/ani13081334. Animals (Basel). 2023. PMID: 37106897 Free PMC article. Review.
-
Campylobacter jejuni in Poultry: Pathogenesis and Control Strategies.Microorganisms. 2022 Oct 28;10(11):2134. doi: 10.3390/microorganisms10112134. Microorganisms. 2022. PMID: 36363726 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

