Exploring the Effect of Differentially Expressed Long Non-coding RNAs Driven by Copy Number Variation on Competing Endogenous RNA Network by Mining Lung Adenocarcinoma Data
- PMID: 33585468
- PMCID: PMC7876300
- DOI: 10.3389/fcell.2020.627436
Exploring the Effect of Differentially Expressed Long Non-coding RNAs Driven by Copy Number Variation on Competing Endogenous RNA Network by Mining Lung Adenocarcinoma Data
Abstract
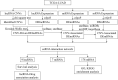
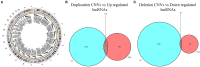
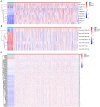
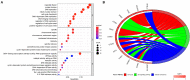
Lung cancer is the first cause of cancer death, and gene copy number variation (CNV) is a vital cause of lung cancer progression. Prognosis prediction of patients followed by medication guidance by detecting CNV of lung cancer is emerging as a promising precise treatment in the future. In this paper, the differences in CNV and gene expression between cancer tissue and normal tissue of lung adenocarcinoma (LUAD) from The Cancer Genome Atlas Lung Adenocarcinoma data set were firstly analyzed, and greater differences were observed. Furthermore, CNV-driven differentially expressed long non-coding RNAs (lncRNAs) were screened out, and then, a competing endogenous RNA (ceRNA) regulatory network related to the gene CNV was established, which involved 9 lncRNAs, seven microRNAs, and 178 downstream messenger RNAs (mRNAs). Pathway enrichment analyses sequentially performed revealed that the downstream mRNAs were mainly enriched in biological pathways related to cell division, DNA repair, and so on, indicating that these mRNAs mainly affected the replication and growth of tumor cells. Besides, the relationship between lncRNAs and drug effects was explored based on previous studies, and it was found that LINC00511 and LINC00942 in the CNV-associated ceRNA network could be used to determine tumor response to drug treatment. As examined, the drugs affected by these two lncRNAs mainly targeted metabolism, target of rapamycin signaling pathway, phosphatidylinositol-3-kinase signaling pathway, epidermal growth factor receptor signaling pathway, and cell cycle. In summary, the present research was devoted to analyzing CNV, lncRNA, mRNA, and microRNA of lung cancer, and nine lncRNAs that could affect the CNV-associated ceRNA network we constructed were identified, two of which are promising in determining tumor response to drug treatment.
Keywords: LUAD; TCGA; ceRNA network; copy number variation; lncRNA.
Copyright © 2021 Hu, Xu, Lu, Zhang, Xu, Xu, Jiang, Zeng, Chen and He.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



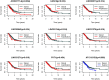
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

