A genomic region associated with protection against severe COVID-19 is inherited from Neandertals
- PMID: 33593941
- PMCID: PMC7936282
- DOI: 10.1073/pnas.2026309118
A genomic region associated with protection against severe COVID-19 is inherited from Neandertals
Abstract
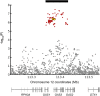
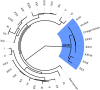
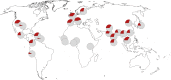
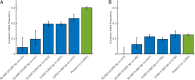
It was recently shown that the major genetic risk factor associated with becoming severely ill with COVID-19 when infected by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is inherited from Neandertals. New, larger genetic association studies now allow additional genetic risk factors to be discovered. Using data from the Genetics of Mortality in Critical Care (GenOMICC) consortium, we show that a haplotype at a region on chromosome 12 associated with requiring intensive care when infected with the virus is inherited from Neandertals. This region encodes proteins that activate enzymes that are important during infections with RNA viruses. In contrast to the previously described Neandertal haplotype that increases the risk for severe COVID-19, this Neandertal haplotype is protective against severe disease. It also differs from the risk haplotype in that it has a more moderate effect and occurs at substantial frequencies in all regions of the world outside Africa. Among ancient human genomes in western Eurasia, the frequency of the protective Neandertal haplotype may have increased between 20,000 and 10,000 y ago and again during the past 1,000 y.
Keywords: COVID-19; Neandertals; OAS1; SARS-CoV-2.
Copyright © 2021 the Author(s). Published by PNAS.
Conflict of interest statement
The authors declare no competing interest.
Figures
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

