Genome-wide identification and functional prediction of long non-coding RNAs in Sprague-Dawley rats during heat stress
- PMID: 33596828
- PMCID: PMC7891137
- DOI: 10.1186/s12864-021-07421-8
Genome-wide identification and functional prediction of long non-coding RNAs in Sprague-Dawley rats during heat stress
Abstract
Background: Heat stress (HS) is a major stress event in the life of an animal, with detrimental upshots in production and health. Long-non-coding RNAs (lncRNAs) play an important role in many biological processes by transcriptional regulation. However, no research has been reported on the characterization and functionality of lncRNAs in heat-stressed rats.
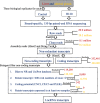
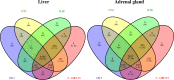
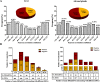
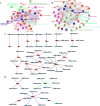
Results: We studied expression levels of lncRNAs in rats during HS, using strand-specific RNA sequencing. Six rats, three in each of Control (22 ± 1 °C) and H120 (42 °C for 120 min) experimental groups, were used to screen for lncRNAs in their liver and adrenal glands. Totally, 4498 and 7627 putative lncRNAs were identified in liver and adrenal glands of the Control and H120 groups, respectively. The majority of lncRNAs were relatively shorter and contained fewer exons than protein-coding transcripts. In total, 482 (174 up-regulated and 308 down-regulated) and 271 (126 up-regulated and 145 down-regulated) differentially-expressed lncRNAs (DElncRNAs, P < 0.05) were identified in the liver and adrenal glands of the Control and H120 groups, respectively. Furthermore, 1274, 121, and 73 target differentially-expressed genes (DEGs) in the liver were predicted to interact with DElncRNAs based on trans-/cis- and sequence similarity regulatory modes. Functional annotation analyses indicated that these DEGs were mostly significantly enriched in insulin signalling, myeloid leukaemia, and glucagon signalling pathways. Similarly, 437, 73 and 41 target DEGs in the adrenal glands were mostly significantly enriched in the cell cycle (trans-prediction) and lysosome pathways (cis-prediction). The DElncRNAs interacting with DEGs that encode heat shock proteins (HSPs) may play an important role in HS response, which include Hsf4, Dnaja1, Dnajb4, Hsph1 and Hspb1 in the liver, and Dnajb13 and Hspb8 in the adrenal glands. The strand-specific RNA sequencing findings were also further verified through RT-qPCR.
Conclusions: This study is the first to provide a detailed characterization and functional analysis of expression levels of lncRNAs in liver and adrenal glands of heat-stressed rats, which provides basis for further studies on the biological functions of lncRNAs under heat stress in rats and other mammalian species.
Keywords: Adrenal glands; DEGs; Heat shock protein; Heat stress response; Liver; LncRNAs.
Conflict of interest statement
The authors declare that they have no conflicts of interest.
Figures


References
-
- Michelozzi P, Accetta G, De Sario M, D'Ippoliti D, Marino C, Baccini M, Biggeri A, Anderson HR, Katsouyanni K, Ballester F, et al. High temperature and hospitalizations for cardiovascular and respiratory causes in 12 European cities. Am J Respir Crit Care Med. 2009;179(5):383–389. doi: 10.1164/rccm.200802-217OC. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

