Opium Poppy Mosaic Virus Has an Xrn-Resistant, Translated Subgenomic RNA and a BTE 3' CITE
- PMID: 33597210
- PMCID: PMC8104115
- DOI: 10.1128/JVI.02109-20
Opium Poppy Mosaic Virus Has an Xrn-Resistant, Translated Subgenomic RNA and a BTE 3' CITE
Abstract
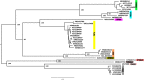
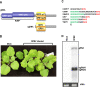
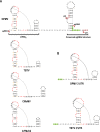
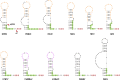
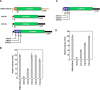
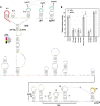
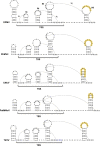
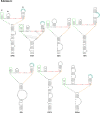
Opium poppy mosaic virus (OPMV) is a recently discovered umbravirus in the family Tombusviridae OPMV has a plus-sense genomic RNA (gRNA) of 4,241 nucleotides (nt) from which replication protein p35 and p35 extension product p98, the RNA-dependent RNA polymerase (RdRp), are expressed. Movement proteins p27 (long distance) and p28 (cell to cell) are expressed from a 1,440-nt subgenomic RNA (sgRNA2). A highly conserved structure was identified just upstream from the sgRNA2 transcription start site in all umbraviruses, which includes a carmovirus consensus sequence, denoting generation by an RdRp-mediated mechanism. OPMV also has a second sgRNA of 1,554 nt (sgRNA1) that starts just downstream of a canonical exoribonuclease-resistant sequence (xrRNAD). sgRNA1 codes for a 30-kDa protein in vitro that is in frame with p28 and cannot be synthesized in other umbraviruses. Eliminating sgRNA1 or truncating the p30 open reading frame (ORF) without affecting p28 substantially reduced accumulation of OPMV gRNA, suggesting a functional role for the protein. The 652-nt 3' untranslated region of OPMV contains two 3' cap-independent translation enhancers (3' CITEs), a T-shaped structure (TSS) near its 3' end, and a Barley yellow dwarf virus-like translation element (BTE) in the central region. Only the BTE is functional in luciferase reporter constructs containing gRNA or sgRNA2 5' sequences in vivo, which differs from how umbravirus 3' CITEs were used in a previous study. Similarly to most 3' CITEs, the OPMV BTE links to the 5' end via a long-distance RNA-RNA interaction. Analysis of 14 BTEs revealed additional conserved sequences and structural features beyond the previously identified 17-nt conserved sequence.IMPORTANCEOpium poppy mosaic virus (OPMV) is an umbravirus in the family Tombusviridae We determined that OPMV accumulates two similarly sized subgenomic RNAs (sgRNAs), with the smaller known to code for proteins expressed from overlapping open reading frames. The slightly larger sgRNA1 has a 5' end just upstream from a previously predicted xrRNAD site, identifying this sgRNA as an unusually long product produced by exoribonuclease trimming. Although four umbraviruses have similar predicted xrRNAD sites, only sgRNA1 of OPMV can code for a protein that is an extension product of umbravirus ORF4. Inability to generate the sgRNA or translate this protein was associated with reduced gRNA accumulation in vivo We also characterized the OPMV BTE structure, a 3' cap-independent translation enhancer (3' CITE). Comparisons of 13 BTEs with the OPMV BTE revealed additional stretches of sequence similarity beyond the 17-nt signature sequence, as well as conserved structural features not previously recognized in these 3' CITEs.
Keywords: 3′ CITE; BTE; Xrn1; cap-independent translation enhancer; exoribonuclease-resistant sites; subgenomic RNA; umbravirus.
Copyright © 2021 American Society for Microbiology.
Figures


Similar articles
-
Conserved Structure Associated with Different 3'CITEs Is Important for Translation of Umbraviruses.Viruses. 2023 Feb 27;15(3):638. doi: 10.3390/v15030638. Viruses. 2023. PMID: 36992347 Free PMC article.
-
trans regulation of cap-independent translation by a viral subgenomic RNA.J Virol. 2006 Oct;80(20):10045-54. doi: 10.1128/JVI.00991-06. J Virol. 2006. PMID: 17005682 Free PMC article.
-
Subgenomic RNA as a riboregulator: negative regulation of RNA replication by Barley yellow dwarf virus subgenomic RNA 2.Virology. 2004 Oct 1;327(2):196-205. doi: 10.1016/j.virol.2004.06.025. Virology. 2004. PMID: 15351207
-
Cis- and trans-regulation of luteovirus gene expression by the 3' end of the viral genome.Virus Res. 2015 Aug 3;206:37-45. doi: 10.1016/j.virusres.2015.03.009. Epub 2015 Apr 6. Virus Res. 2015. PMID: 25858272 Free PMC article. Review.
-
Host-like RNA Elements Regulate Virus Translation.Viruses. 2024 Mar 20;16(3):468. doi: 10.3390/v16030468. Viruses. 2024. PMID: 38543832 Free PMC article. Review.
Cited by
-
Identification of Novel 5' and 3' Translation Enhancers in Umbravirus-Like Coat Protein-Deficient RNA Replicons.J Virol. 2022 Apr 13;96(7):e0173621. doi: 10.1128/jvi.01736-21. Epub 2022 Mar 17. J Virol. 2022. PMID: 35297668 Free PMC article.
-
Umbravirus-like RNA viruses are capable of independent systemic plant infection in the absence of encoded movement proteins.PLoS Biol. 2024 Apr 25;22(4):e3002600. doi: 10.1371/journal.pbio.3002600. eCollection 2024 Apr. PLoS Biol. 2024. PMID: 38662792 Free PMC article.
-
Conserved Structure Associated with Different 3'CITEs Is Important for Translation of Umbraviruses.Viruses. 2023 Feb 27;15(3):638. doi: 10.3390/v15030638. Viruses. 2023. PMID: 36992347 Free PMC article.
-
RNA Structure Protects the 5' End of an Uncapped Tombusvirus RNA Genome from Xrn Digestion.J Virol. 2021 Sep 27;95(20):e0103421. doi: 10.1128/JVI.01034-21. Epub 2021 Aug 4. J Virol. 2021. PMID: 34346764 Free PMC article.
-
Translation of Plant RNA Viruses.Viruses. 2021 Dec 13;13(12):2499. doi: 10.3390/v13122499. Viruses. 2021. PMID: 34960768 Free PMC article. Review.
References
-
- Kim SH, MacFarlane S, Kalinina NO, Rakitina DV, Ryabov EV, Gillespie T, Haupt S, Brown JWS, Taliansky M. 2007. Interaction of a plant virus-encoded protein with the major nucleolar protein fibrillarin is required for systemic virus infection. Proc Natl Acad Sci U S A 104:11115–11120. 10.1073/pnas.0704632104. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

