Assessment of S1-, S2-, and NCP-Specific IgM, IgA, and IgG Antibody Kinetics in Acute SARS-CoV-2 Infection by a Microarray and Twelve Other Immunoassays
- PMID: 33602698
- PMCID: PMC8091850
- DOI: 10.1128/JCM.02890-20
Assessment of S1-, S2-, and NCP-Specific IgM, IgA, and IgG Antibody Kinetics in Acute SARS-CoV-2 Infection by a Microarray and Twelve Other Immunoassays
Abstract
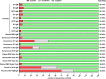
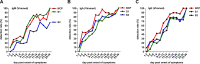
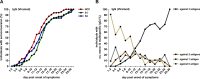
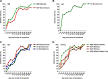
In this study, we comprehensively analyzed multispecific antibody kinetics of different immunoglobulins in hospitalized patients with acute severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection. Three hundred fifty-four blood samples longitudinally obtained from 81 IgG-seroconverting progressed coronavirus disease 2019 (CoVID-19) patients were quantified for spike 1 (S1), S2, and nucleocapsid protein (NCP)-specific IgM, IgA, IgG, and total Ig antibodies using a microarray, 11 different enzyme-linked immunosorbent assays (ELISAs)/chemiluminescence immunoassays (CLIAs), and 1 rapid test by seven manufacturers. The assays' specificity was assessed in 130 non-CoVID-19 pneumonia patients. Using the microarray, NCP-specific IgA and IgG antibodies continuously displayed higher detection rates during acute CoVID-19 than S1- and S2-specific ones. S1-specific IgG antibodies, however, reached higher peak values. Until the 26th day post-symptom onset, all patients developed IgG responses against S1, S2, and NCP. Although detection rates by ELISAs/CLIAs generally resembled those of the microarray, corresponding to the target antigen, sensitivities and specificities varied among all tests. Notably, patients with more severe CoVID-19 displayed higher IgG and IgA levels, but this difference was mainly observed with S1-specific immunoassays. In patients with high SARS-CoV-2 levels in the lower respiratory tract, we observed high detection rates of IgG and total Ig immunoassays with a particular rise of S1-specific IgG antibodies when viral concentrations in the tracheal aspirate subsequently declined over time. In summary, our study demonstrates that differences in sensitivity among commercial immunoassays during acute SARS-CoV-2 infection are only partly related to the target antigen. Importantly, our data indicate that NCP-specific IgA and IgG antibodies are detected earlier, while higher S1-specific IgA antibody levels occur in severely ill patients.
Keywords: ELISA; IgA; SARS; SARS-CoV-2; antibodies; coronavirus; immunoassay; microarray.
Copyright © 2021 American Society for Microbiology.
Figures

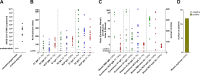

References
-
- Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, Zhao X, Huang B, Shi W, Lu R, Niu P, Zhan F, Ma X, Wang D, Xu W, Wu G, Gao GF, Tan W, China Novel Coronavirus Investigating and Research Team . 2020. A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med 382:727–733. 10.1056/NEJMoa2001017. - DOI - PMC - PubMed
-
- Corman VM, Landt O, Kaiser M, Molenkamp R, Meijer A, Chu DKW, Bleicker T, Brunink S, Schneider J, Schmidt ML, Mulders D, Haagmans BL, van der Veer B, van den Brink S, Wijsman L, Goderski G, Romette JL, Ellis J, Zambon M, Peiris M, Goossens H, Reusken C, Koopmans MPG, Drosten C. 2020. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill 25:2000045. 10.2807/1560-7917.ES.2020.25.3.2000045. - DOI - PMC - PubMed
-
- Zhao J, Yuan Q, Wang H, Liu W, Liao X, Su Y, Wang X, Yuan J, Li T, Li J, Qian S, Hong C, Wang F, Liu Y, Wang Z, He Q, Li Z, He B, Zhang T, Fu Y, Ge S, Liu L, Zhang J, Xia N, Zhang Z. 2020. Antibody responses to SARS-CoV-2 in patients of novel coronavirus disease 2019. Clin Infect Dis 71:2027–2034. 10.1093/cid/ciaa344. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

