EARLY BUD-BREAK 1 and EARLY BUD-BREAK 3 control resumption of poplar growth after winter dormancy
- PMID: 33602938
- PMCID: PMC7893051
- DOI: 10.1038/s41467-021-21449-0
EARLY BUD-BREAK 1 and EARLY BUD-BREAK 3 control resumption of poplar growth after winter dormancy
Abstract
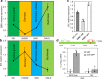
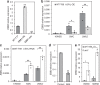
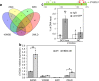
Bud-break is an economically and environmentally important process in trees and shrubs from boreal and temperate latitudes, but its molecular mechanisms are poorly understood. Here, we show that two previously reported transcription factors, EARLY BUD BREAK 1 (EBB1) and SHORT VEGETATIVE PHASE-Like (SVL) directly interact to control bud-break. EBB1 is a positive regulator of bud-break, whereas SVL is a negative regulator of bud-break. EBB1 directly and negatively regulates SVL expression. We further report the identification and characterization of the EBB3 gene. EBB3 is a temperature-responsive, epigenetically-regulated, positive regulator of bud-break that provides a direct link to activation of the cell cycle during bud-break. EBB3 is an AP2/ERF transcription factor that positively and directly regulates CYCLIND3.1 gene. Our results reveal the architecture of a putative regulatory module that links temperature-mediated control of bud-break with activation of cell cycle.
Conflict of interest statement
The authors declare no competing interests.
Figures




References
-
- Howe GT, Hackett WP, Furnier GR, Klevorn RE. Photoperiodic responses of a northern and southern ecotype of black cottonwood. Physiologia Plant. 1995;93:695–708. doi: 10.1111/j.1399-3054.1995.tb05119.x. - DOI
-
- Jeknić Z, Chen THH. Changes in protein profiles of poplar tissues during the induction of bud dormancy by short-day photoperiods1. Plant Cell Physiol. 1999;40:25–35. doi: 10.1093/oxfordjournals.pcp.a029471. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

