Identifying Differentially Expressed Genes of Zero Inflated Single Cell RNA Sequencing Data Using Mixed Model Score Tests
- PMID: 33613638
- PMCID: PMC7894898
- DOI: 10.3389/fgene.2021.616686
Identifying Differentially Expressed Genes of Zero Inflated Single Cell RNA Sequencing Data Using Mixed Model Score Tests
Abstract
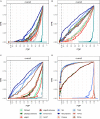
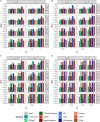
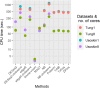
Single cell RNA sequencing (scRNA-seq) allows quantitative measurement and comparison of gene expression at the resolution of single cells. Ignoring the batch effects and zero inflation of scRNA-seq data, many proposed differentially expressed (DE) methods might generate bias. We propose a method, single cell mixed model score tests (scMMSTs), to efficiently identify DE genes of scRNA-seq data with batch effects using the generalized linear mixed model (GLMM). scMMSTs treat the batch effect as a random effect. For zero inflation, scMMSTs use a weighting strategy to calculate observational weights for counts independently under zero-inflated and zero-truncated distributions. Counts data with calculated weights were subsequently analyzed using weighted GLMMs. The theoretical null distributions of the score statistics were constructed by mixed Chi-square distributions. Intensive simulations and two real datasets were used to compare edgeR-zinbwave, DESeq2-zinbwave, and scMMSTs. Our study demonstrates that scMMSTs, as supplement to standard methods, are advantageous to define DE genes of zero-inflated scRNA-seq data with batch effects.
Keywords: differential expression analyses; generalized linear mixed model; observational weights; score test; single cell RNA sequencing; zero inflation.
Copyright © 2021 He, Pan, Shao and Wang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Observation weights unlock bulk RNA-seq tools for zero inflation and single-cell applications.Genome Biol. 2018 Feb 26;19(1):24. doi: 10.1186/s13059-018-1406-4. Genome Biol. 2018. PMID: 29478411 Free PMC article.
-
ZERO-INFLATED QUANTILE RANK-SCORE BASED TEST (ZIQRANK) WITH APPLICATION TO SCRNA-SEQ DIFFERENTIAL GENE EXPRESSION ANALYSIS.Ann Appl Stat. 2021 Dec;15(4):1673-1696. doi: 10.1214/21-aoas1442. Epub 2021 Dec 21. Ann Appl Stat. 2021. PMID: 35116085 Free PMC article.
-
Differential expression of single-cell RNA-seq data using Tweedie models.Stat Med. 2022 Aug 15;41(18):3492-3510. doi: 10.1002/sim.9430. Epub 2022 Jun 2. Stat Med. 2022. PMID: 35656596 Free PMC article.
-
Bayesian gamma-negative binomial modeling of single-cell RNA sequencing data.BMC Genomics. 2020 Sep 9;21(Suppl 9):585. doi: 10.1186/s12864-020-06938-8. BMC Genomics. 2020. PMID: 32900358 Free PMC article.
-
Naught all zeros in sequence count data are the same.Comput Struct Biotechnol J. 2020 Sep 28;18:2789-2798. doi: 10.1016/j.csbj.2020.09.014. eCollection 2020. Comput Struct Biotechnol J. 2020. PMID: 33101615 Free PMC article. Review.
Cited by
-
Challenges and best practices in omics benchmarking.Nat Rev Genet. 2024 May;25(5):326-339. doi: 10.1038/s41576-023-00679-6. Epub 2024 Jan 12. Nat Rev Genet. 2024. PMID: 38216661 Review.
-
Differential Expression Analysis of Single-Cell RNA-Seq Data: Current Statistical Approaches and Outstanding Challenges.Entropy (Basel). 2022 Jul 18;24(7):995. doi: 10.3390/e24070995. Entropy (Basel). 2022. PMID: 35885218 Free PMC article. Review.
-
Leveraging gene correlations in single cell transcriptomic data.BMC Bioinformatics. 2024 Sep 18;25(1):305. doi: 10.1186/s12859-024-05926-z. BMC Bioinformatics. 2024. PMID: 39294560 Free PMC article.
-
Leveraging gene correlations in single cell transcriptomic data.bioRxiv [Preprint]. 2023 Nov 1:2023.03.14.532643. doi: 10.1101/2023.03.14.532643. bioRxiv. 2023. Update in: BMC Bioinformatics. 2024 Sep 18;25(1):305. doi: 10.1186/s12859-024-05926-z. PMID: 36993765 Free PMC article. Updated. Preprint.
References
-
- Benjamini Y., Hochberg Y. (1995). Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B-Methodol. 57 289–300. 10.1111/j.2517-6161.1995.tb02031.x - DOI
-
- Böhning D., Dietz E., Schlattmann P., Mendonça L., Kirchner U. (1999). The zero-inflated Poisson model and the decayed, missing and filled teeth index in dental epidemiology. J. R. Stat. Soc. Ser. A 162 195–209. 10.1111/1467-985X.00130 - DOI
-
- Breslow N. E., Clayton D. G. (1993). Approximate inference in generalized linear mixed models. J. Am. Stat. Assoc. 88 9–25. 10.2307/2290687 - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources

