Weaning Induces Stress-Dependent DNA Methylation and Transcriptional Changes in Piglet PBMCs
- PMID: 33613645
- PMCID: PMC7893110
- DOI: 10.3389/fgene.2021.633564
Weaning Induces Stress-Dependent DNA Methylation and Transcriptional Changes in Piglet PBMCs
Abstract
Changes to the epigenome, including those to DNA methylation, have been proposed as mechanisms by which stress can induce long-term physiological changes in livestock species. Pig weaning is associated with dietary and social stress, both of which elicit an immune response and changes to the hypothalamic-pituitary-adrenal (HPA) axis. While differential methylation following stress has been assessed in model organisms, it remains poorly understood how the pig methylome is altered by stressors in production settings. We quantified changes in CpG methylation and transcript abundance in piglet peripheral blood mononuclear cells (PBMCs) following weaning and also assessed differential patterns in pigs exhibiting high and low stress response as measured by cortisol concentration and lesion scores. Blood was collected from nine gilt piglets 24 h before and after weaning, and whole-genome bisulfite sequencing (WGBS) and RNA-sequencing were performed on six and nine animals, respectively, at both time points. We identified 2,674 differentially methylated regions (DMRs) that were enriched within promoters of genes associated with lymphocyte stimulation and transcriptional regulation. Stress groups displayed unique differential methylation and expression patterns associated with activation and suppression of T cell immunity in low and high stress animals, respectively. Differential methylation was strongly associated with differential expression; specifically, upregulated genes were enriched among hypomethylated genes. We observed post-weaning hypermethylation of the glucocorticoid receptor (NR3C1) promoter and a significant decrease in NR3C1 expression (n = 9, p = 6.1 × 10-3). Our results indicate that weaning-associated stress elicits genome-wide methylation changes associated with differential gene expression, reduced T cell activation, and an altered HPA axis response.
Keywords: DNA methylation; NR3C1; epigenetics; gene expression; pig; weaning.
Copyright © 2021 Corbett, Luttman, Wurtz, Siegford, Raney, Ford and Ernst.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
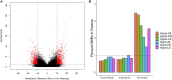

References
-
- Ashcraft K. A., Hunzeker J., Bonneau R. H. (2008). Psychological stress impairs the local CD8+ T cell response to mucosal HSV-1 infection and allows for increased pathogenicity via a glucocorticoid receptor-mediated mechanism. Psychoneuroendocrinology 33 951–963. 10.1016/j.psyneuen.2008.04.010 - DOI - PMC - PubMed
-
- Baker E. C., Cilkiz K. Z., Riggs P. K., Littlejohn B. P., Long C. R., Welsh T. H., et al. (2020). Effect of prenatal transportation stress on DNA methylation in Brahman heifers. Livest. Sci. 240:104116 10.1016/j.livsci.2020.104116 - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources

