Identification of novel avian and mammalian deltaviruses provides new insights into deltavirus evolution
- PMID: 33614159
- PMCID: PMC7882216
- DOI: 10.1093/ve/veab003
Identification of novel avian and mammalian deltaviruses provides new insights into deltavirus evolution
Abstract
Hepatitis delta virus (HDV) is a satellite virus that requires hepadnavirus envelope proteins for its transmission. Although recent studies identified HDV-related deltaviruses in certain animals, the evolution of deltaviruses, such as the origin of HDV and the mechanism of its coevolution with its helper viruses, is unknown, mainly because of the phylogenetic gaps among deltaviruses. Here, we identified novel deltaviruses of passerine birds, woodchucks, and white-tailed deer by extensive database searches and molecular surveillance. Phylogenetic and molecular epidemiological analyses suggest that HDV originated from mammalian deltaviruses and the past interspecies transmission of mammalian and passerine deltaviruses. Further, metaviromic and experimental analyses suggest that the satellite-helper relationship between HDV and hepadnavirus was established after the divergence of the HDV lineage from non-HDV mammalian deltaviruses. Our findings enhance our understanding of deltavirus evolution, diversity, and transmission, indicating the importance of further surveillance for deltaviruses.
Keywords: deltavirus; inter-species transmission; virome.
© The Author(s) 2021. Published by Oxford University Press.
Figures





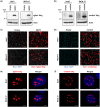

References
-
- Alvarado-Mora M. V. et al. (2011) ‘Dynamics of Hepatitis D (Delta) Virus Genotype 3 in the Amazon Region of South America’, Infection, Genetics and Evolution, 11: 1462–8. - PubMed
-
- Bergmann K. F., Gerin J. L. (1986) ‘ Antigens of Hepatitis Delta Virus in the Liver and Serum of Humans and Animals’, Journal of Infectious Diseases, 154: 702–6. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

