Development of a new genetic reference material system based on Saccharomyces cerevisiae cells
- PMID: 33614823
- PMCID: PMC7868937
- DOI: 10.1016/j.omtm.2021.01.004
Development of a new genetic reference material system based on Saccharomyces cerevisiae cells
Abstract
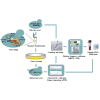
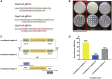
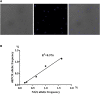
As an important quality control link of molecular diagnosis, genetic reference materials (RMs) are widely used in various gene detection platforms such as mutation detection, gene quantification, and second generation sequencing. However, contamination, construction, and storage of existing genetic RMs still remain challenges. Here, we established a new genetic RM system based on Saccharomyces cerevisiae. We chose the non-small cell lung cancer (NSCLC) mutation hotspots in Kirsten rat sarcoma viral oncogene (KRAS) and epidermal growth factor receptor (EGFR), using clustered regularly interspaced short palindromic repeats and CRISPR-associated protein (CRISPR-Cas9) system-mediated gene editing technology, combined with the high homologous recombination efficiency of Saccharomyces cerevisiae. A single copy of the target gene was inserted into the yeast genome, and the inserted target gene was stably inherited with the passage of yeast cells. The copy number calculation for the target gene can replays by cell counting. The RM system was evaluated by sequence, copy number, stability, and homogeneity. In summary, the recombinant yeast cell line has ease of construction and screening, stable genetic characteristics, accurate copy number calculation, and convenient culture and preservation. Our findings may provide new ideas and directions for the research and industrialization of genetic RMs.
Keywords: CRISPR-Cas9; EGFR; KRAS; Saccharomyces cerevisiae; genetic reference materials; homologous recombination.
© 2021 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
Homology-integrated CRISPR-Cas (HI-CRISPR) system for one-step multigene disruption in Saccharomyces cerevisiae.ACS Synth Biol. 2015 May 15;4(5):585-94. doi: 10.1021/sb500255k. Epub 2014 Sep 19. ACS Synth Biol. 2015. PMID: 25207793
-
Two Distinct Approaches for CRISPR-Cas9-Mediated Gene Editing in Cryptococcus neoformans and Related Species.mSphere. 2018 Jun 13;3(3):e00208-18. doi: 10.1128/mSphereDirect.00208-18. Print 2018 Jun 27. mSphere. 2018. PMID: 29898980 Free PMC article.
-
Gene Therapy with CRISPR/Cas9 Coming to Age for HIV Cure.AIDS Rev. 2017 Oct-Dec;19(3):167-172. AIDS Rev. 2017. PMID: 29019352
-
CRISPR system in the yeast Saccharomyces cerevisiae and its application in the bioproduction of useful chemicals.World J Microbiol Biotechnol. 2019 Jul 6;35(7):111. doi: 10.1007/s11274-019-2688-8. World J Microbiol Biotechnol. 2019. PMID: 31280424 Review.
-
Gene Editing in Clinical Practice: Where are We?Indian J Clin Biochem. 2019 Jan;34(1):19-25. doi: 10.1007/s12291-018-0804-4. Epub 2019 Jan 1. Indian J Clin Biochem. 2019. PMID: 30728669 Free PMC article. Review.
References
-
- Hawkins J.R., Hawkins M., Boyle J., Gray E., Matejtschuk P., Metcalfe P. Genetic reference materials and their application to haematology. Biologicals. 2010;38:467–473. - PubMed
-
- Scharf H., Bremser W. Certification of a reference material for determination of total cyanide in soil to support implementation of the International Standard ISO 11262:2011. Anal. Bioanal. Chem. 2015;407:3219–3223. - PubMed
-
- Grimalt S., Harbeck S., Shegunova P., Seghers J., Sejerøe-Olsen B., Emteborg H., Dabrio M. Development of a new cucumber reference material for pesticide residue analysis: feasibility study for material processing, homogeneity and stability assessment. Anal. Bioanal. Chem. 2015;407:3083–3091. - PMC - PubMed
-
- Levin B.C., Richie K.L., Jakupciak J.P. Advances in Huntington’s disease diagnostics: development of a standard reference material. Expert Rev. Mol. Diagn. 2006;6:587–596. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

