c-Myb interferes with inflammatory IL1α-NF-κB pathway in breast cancer cells
- PMID: 33621853
- PMCID: PMC7905261
- DOI: 10.1016/j.neo.2021.01.002
c-Myb interferes with inflammatory IL1α-NF-κB pathway in breast cancer cells
Abstract
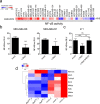
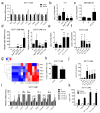
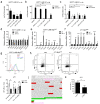
The transcription factor c-Myb can be involved in the activation of many genes with protumorigenic function; however, its role in breast cancer (BC) development is still under discussion. c-Myb is considered as a tumor-promoting factor in the early phases of BC, on the other hand, its expression in BC patients relates to a good prognosis. Previously, we have shown that c-Myb controls the capacity of BC cells to form spontaneous lung metastasis. Reduced seeding of BC cells to the lungs is linked to high expression of c-Myb and a decline in expression of a specific set of inflammatory genes. Here, we unraveled a c-Myb-IL1α-NF-κB signaling axis that takes place in tumor cells. We report that an overexpression of c-Myb interfered with the activity of NF-κB in several BC cell lines. We identified IL1α to be essential for this interference since it was abrogated in the IL1α-deficient cells. Overexpression of IL1α, as well as addition of recombinant IL1α protein, activated NF-κB signaling and restored expression of the inflammatory signature genes suppressed by c-Myb. The endogenous levels of c-Myb negatively correlated with IL1α on both transcriptional and protein levels across BC cell lines. We concluded that inhibition of IL1α expression by c-Myb reduces NF-κB activity and disconnects the inflammatory circuit, a potentially targetable mechanism to mimic the antimetastatic effect of c-Myb with therapeutic perspective.
Keywords: Breast cancer; IL1α; Inflammation; NF-κB; Transactivation; c-Myb.
Copyright © 2021 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of competing interest The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures

Similar articles
-
Transcription factor c-Myb inhibits breast cancer lung metastasis by suppression of tumor cell seeding.Oncogene. 2018 Feb 22;37(8):1020-1030. doi: 10.1038/onc.2017.392. Epub 2017 Oct 30. Oncogene. 2018. PMID: 29084208 Free PMC article.
-
Human T-cell lymphotropic virus type 1 Tax represses c-Myb-dependent transcription through activation of the NF-kappaB pathway and modulation of coactivator usage.Mol Cell Biol. 2001 Nov;21(21):7391-402. doi: 10.1128/MCB.21.21.7391-7402.2001. Mol Cell Biol. 2001. PMID: 11585920 Free PMC article.
-
HER2 Overexpression Triggers an IL1α Proinflammatory Circuit to Drive Tumorigenesis and Promote Chemotherapy Resistance.Cancer Res. 2018 Apr 15;78(8):2040-2051. doi: 10.1158/0008-5472.CAN-17-2761. Epub 2018 Jan 30. Cancer Res. 2018. PMID: 29382706 Free PMC article.
-
New Insights on COX-2 in Chronic Inflammation Driving Breast Cancer Growth and Metastasis.J Mammary Gland Biol Neoplasia. 2015 Dec;20(3-4):109-19. doi: 10.1007/s10911-015-9333-4. Epub 2015 Jul 21. J Mammary Gland Biol Neoplasia. 2015. PMID: 26193871 Review.
-
The role of NF-κB in breast cancer initiation, growth, metastasis, and resistance to chemotherapy.Biomed Pharmacother. 2023 Jul;163:114822. doi: 10.1016/j.biopha.2023.114822. Epub 2023 May 3. Biomed Pharmacother. 2023. PMID: 37146418 Review.
Cited by
-
Integrated enhancer regulatory network by enhancer-promoter looping in gastric cancer.NAR Cancer. 2024 May 8;6(2):zcae020. doi: 10.1093/narcan/zcae020. eCollection 2024 Jun. NAR Cancer. 2024. PMID: 38720882 Free PMC article.
-
Transcription factor c-Myb: novel prognostic factor in osteosarcoma.Clin Exp Metastasis. 2022 Apr;39(2):375-390. doi: 10.1007/s10585-021-10145-4. Epub 2022 Jan 7. Clin Exp Metastasis. 2022. PMID: 34994868
-
Inflammatory Breast Cancer: The Cytokinome of Post-Mastectomy Wound Fluid Augments Proliferation, Invasion, and Stem Cell Markers.Curr Issues Mol Biol. 2022 Jun 17;44(6):2730-2744. doi: 10.3390/cimb44060187. Curr Issues Mol Biol. 2022. PMID: 35735628 Free PMC article.
-
CRP, IL-1α, IL-1β, and IL-6 levels and the risk of breast cancer: a two-sample Mendelian randomization study.Sci Rep. 2024 Jan 23;14(1):1982. doi: 10.1038/s41598-024-52080-w. Sci Rep. 2024. PMID: 38263420 Free PMC article.
-
Aged Breast Extracellular Matrix Drives Mammary Epithelial Cells to an Invasive and Cancer-Like Phenotype.Adv Sci (Weinh). 2021 Nov;8(22):e2100128. doi: 10.1002/advs.202100128. Epub 2021 Oct 7. Adv Sci (Weinh). 2021. PMID: 34617419 Free PMC article.
References
-
- Calabretta B, Sims RB, Valtieri M, Caracciolo D, Szczylik C, Venturelli D, Ratajczak M, Beran M, Gewirtz AM. Normal and leukemic hematopoietic cells manifest differential sensitivity to inhibitory effects of c-myb antisense oligodeoxynucleotides: an in vitro study relevant to bone marrow purging. Proc Natl Acad Sci U S A. 1991;88:2351–2355. doi: 10.1073/pnas.88.6.2351. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

