Beryllium-specific CD4+ T cells induced by chemokine neoantigens perpetuate inflammation
- PMID: 33630763
- PMCID: PMC8087207
- DOI: 10.1172/JCI144864
Beryllium-specific CD4+ T cells induced by chemokine neoantigens perpetuate inflammation
Abstract
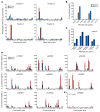
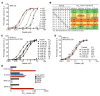
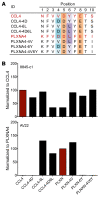
Discovering dominant epitopes for T cells, particularly CD4+ T cells, in human immune-mediated diseases remains a significant challenge. Here, we used bronchoalveolar lavage (BAL) cells from HLA-DP2-expressing patients with chronic beryllium disease (CBD), a debilitating granulomatous lung disorder characterized by accumulations of beryllium-specific (Be-specific) CD4+ T cells in the lung. We discovered lung-resident CD4+ T cells that expressed a disease-specific public CDR3β T cell receptor motif and were specific to Be-modified self-peptides derived from C-C motif ligand 4 (CCL4) and CCL3. HLA-DP2-CCL/Be tetramer staining confirmed that these chemokine-derived peptides represented major antigenic targets in CBD. Furthermore, Be induced CCL3 and CCL4 secretion in the lungs of mice and humans. In a murine model of CBD, the addition of LPS to Be oxide exposure enhanced CCL4 and CCL3 secretion in the lung and significantly increased the number and percentage of CD4+ T cells specific for the HLA-DP2-CCL/Be epitope. Thus, we demonstrate a direct link between Be-induced innate production of chemokines and the development of a robust adaptive immune response to those same chemokines presented as Be-modified self-peptides, creating a cycle of innate and adaptive immune activation.
Keywords: Adaptive immunity; Pulmonology; T cell receptor; T cells.
Conflict of interest statement
Figures





Comment in
-
Detection of autoreactive CD4+ T cells by MHC class II multimers in HLA-linked human autoimmune diseases.J Clin Invest. 2021 May 3;131(9):e148674. doi: 10.1172/JCI148674. J Clin Invest. 2021. PMID: 33938450 Free PMC article.
Similar articles
-
Metal-specific CD4+ T-cell responses induced by beryllium exposure in HLA-DP2 transgenic mice.Mucosal Immunol. 2016 Jan;9(1):218-28. doi: 10.1038/mi.2015.54. Epub 2015 Jul 1. Mucosal Immunol. 2016. PMID: 26129650 Free PMC article.
-
Identification of multiple public TCR repertoires in chronic beryllium disease.J Immunol. 2014 May 15;192(10):4571-80. doi: 10.4049/jimmunol.1400007. Epub 2014 Apr 9. J Immunol. 2014. PMID: 24719461 Free PMC article. Clinical Trial.
-
Regulatory T cells modulate granulomatous inflammation in an HLA-DP2 transgenic murine model of beryllium-induced disease.Proc Natl Acad Sci U S A. 2014 Jun 10;111(23):8553-8. doi: 10.1073/pnas.1408048111. Epub 2014 May 27. Proc Natl Acad Sci U S A. 2014. PMID: 24912188 Free PMC article.
-
Adaptive Immunity in Pulmonary Sarcoidosis and Chronic Beryllium Disease.Front Immunol. 2020 Mar 18;11:474. doi: 10.3389/fimmu.2020.00474. eCollection 2020. Front Immunol. 2020. PMID: 32256501 Free PMC article. Review.
-
Beryllium-Induced Hypersensitivity: Genetic Susceptibility and Neoantigen Generation.J Immunol. 2016 Jan 1;196(1):22-7. doi: 10.4049/jimmunol.1502011. J Immunol. 2016. PMID: 26685315 Free PMC article. Review.
Cited by
-
Protective role of tissue-resident Tregs in a murine model of beryllium-induced disease.JCI Insight. 2022 Aug 22;7(16):e156098. doi: 10.1172/jci.insight.156098. JCI Insight. 2022. PMID: 35819849 Free PMC article.
-
CD4+ T cells in the lungs of acute sarcoidosis patients recognize an Aspergillus nidulans epitope.J Exp Med. 2021 Oct 4;218(10):e20210785. doi: 10.1084/jem.20210785. Epub 2021 Aug 19. J Exp Med. 2021. PMID: 34410304 Free PMC article.
-
Innate and Adaptive Immunity in Noninfectious Granulomatous Lung Disease.J Immunol. 2022 Apr 15;208(8):1835-1843. doi: 10.4049/jimmunol.2101159. J Immunol. 2022. PMID: 35418504 Free PMC article. Review.
-
Detection of autoreactive CD4+ T cells by MHC class II multimers in HLA-linked human autoimmune diseases.J Clin Invest. 2021 May 3;131(9):e148674. doi: 10.1172/JCI148674. J Clin Invest. 2021. PMID: 33938450 Free PMC article.
-
Measuring anti-islet autoimmunity in mouse and human by profiling peripheral blood antigen-specific CD4 T cells.Sci Transl Med. 2023 Jul 5;15(703):eade3614. doi: 10.1126/scitranslmed.ade3614. Epub 2023 Jul 5. Sci Transl Med. 2023. PMID: 37406136 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

