Target-induced clustering activates Trim-Away of pathogens and proteins
- PMID: 33633400
- PMCID: PMC7611929
- DOI: 10.1038/s41594-021-00560-2
Target-induced clustering activates Trim-Away of pathogens and proteins
Abstract
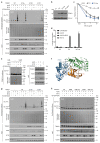
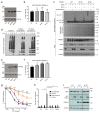
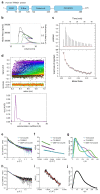
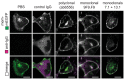
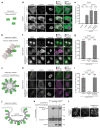
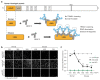
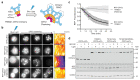
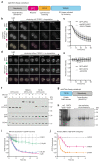
Trim-Away is a recently developed technology that exploits off-the-shelf antibodies and the RING E3 ligase and cytosolic antibody receptor TRIM21 to carry out rapid protein depletion. How TRIM21 is catalytically activated upon target engagement, either during its normal immune function or when repurposed for targeted protein degradation, is unknown. Here we show that a mechanism of target-induced clustering triggers intermolecular dimerization of the RING domain to switch on the ubiquitination activity of TRIM21 and induce virus neutralization or drive Trim-Away. We harness this mechanism for selective degradation of disease-causing huntingtin protein containing long polyglutamine tracts and expand the Trim-Away toolbox with highly active TRIM21-nanobody chimeras that can also be controlled optogenetically. This work provides a mechanism for cellular activation of TRIM RING ligases and has implications for targeted protein degradation technologies.
Conflict of interest statement
The authors declare no competing interests.
Figures



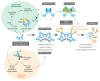
References
-
- Zheng N, Shabek N. Ubiquitin Ligases: Structure, Function, and Regulation. Annu Rev Biochem. 2017;86:129–157. - PubMed
-
- Schapira M, Calabrese MF, Bullock AN, Crews CM. Targeted protein degradation: expanding the toolbox. Nat Rev Drug Discov. 2019;18:949–963. - PubMed
-
- Verma R, Mohl D, Deshaies RJ. Harnessing the Power of Proteolysis for Targeted Protein Inactivation. Mol Cell. 2020;77:446–460. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

