Reductive Transformation of Fe(III) (oxyhydr)Oxides by Mesophilic Homoacetogens in the Genus Sporomusa
- PMID: 33633701
- PMCID: PMC7901989
- DOI: 10.3389/fmicb.2021.600808
Reductive Transformation of Fe(III) (oxyhydr)Oxides by Mesophilic Homoacetogens in the Genus Sporomusa
Abstract
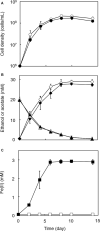
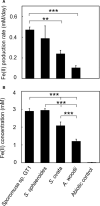
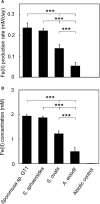
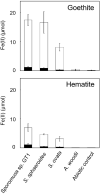
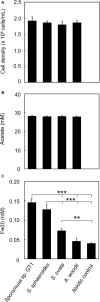
Microbial reduction of iron contributes to the dissolution and transformation of iron-containing minerals in nature. Diverse groups of homoacetogenic bacteria (homoacetogens) have been reported to reduce insoluble Fe(III) oxides, such as hydrous ferric oxide (HFO), an Fe(III) mineral commonly found in soils and sediments. Several members of genus Sporomusa reportedly oxidize Fe(0), indicating the presence of an extracellular electron-uptake mechanism. However, the ability of the genus to reduce insoluble Fe(III) oxides is limited, and the underlying reduction mechanism remains to be elucidated. In this study, the HFO reduction ability of three Sporomusa spp. (Sporomusa sp. strain GT1, Sporomusa sphaeroides, and Sporomusa ovata) and a homoacetogen of a different genus (Acetobacterium woodii) were assayed under organotrophic (ethanol) and lithotrophic (H2 + CO2) conditions without a chelator or reducing reagent. All tested homoacetogens showed acetogenic growth and concomitant reduction of HFO under both organotrophic and lithotrophic conditions. Analysis of the growth stoichiometry showed that Fe(III) reduction does not support direct energy conservation, thereby indicating that Fe(III) reduction is a side reaction of acetogenesis to dissipate the excess reducing power. HFO was reduced to a soluble Fe(II) form by microbial activity. In addition, we observed that strain GT1, S. sphaeroides, and S. ovata reduced crystalline Fe(III) oxides, and HFO was reductively transformed into magnetite (Fe3O4) under phosphate-limiting conditions. Separation of HFO by a dialysis membrane still permitted Fe(II) production, although the reduction rate was decreased, suggesting that Fe(III) reduction is at least partially mediated by soluble redox compound(s) secreted from the cells. Finally, culture experiments and comparative genomic analysis suggested that electron transfer by flavins and multiheme c-type cytochrome were not directly correlated with Fe(III) reduction activity. This study reveals the capability of Sporomusa spp. in the reductive transformation of iron mineral and indicates the potential involvement of these organisms in iron and other mineral cycles in nature.
Keywords: element cycle; extracellular electron transfer; homoacetogenic bacteria; iron reduction; mineral transformation.
Copyright © 2021 Igarashi and Kato.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



References
-
- Aneksampant A., Nakashima K., Kawasaki S. (2020). Microbial leaching of iron from hematite: direct or indirect elution. Mater. Trans. 61 396–401. 10.2320/matertrans.m-m2019860 - DOI
-
- Balch W. E., Schoberth S., Tanner R. S., Wolfe R. S. (1977). Acetobacterium, a new genus of hydrogen-oxidizing, carbon dioxide-reducing, anaerobic bacteria. Int. J. Syst. Bacteriol. 27 355–361. 10.1099/00207713-27-4-355 - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

