Algorithmic Annotation of Functional Roles for Components of 3,044 Human Molecular Pathways
- PMID: 33633781
- PMCID: PMC7900570
- DOI: 10.3389/fgene.2021.617059
Algorithmic Annotation of Functional Roles for Components of 3,044 Human Molecular Pathways
Abstract
Current methods of high-throughput molecular and genomic analyses enabled to reconstruct thousands of human molecular pathways. Knowledge of molecular pathways structure and architecture taken along with the gene expression data can help interrogating the pathway activation levels (PALs) using different bioinformatic algorithms. In turn, the pathway activation profiles can characterize molecular processes, which are differentially regulated and give numeric characteristics of the extent of their activation or inhibition. However, different pathway nodes may have different functions toward overall pathway regulation, and calculation of PAL requires knowledge of molecular function of every node in the pathway in terms of its activator or inhibitory role. Thus, high-throughput annotation of functional roles of pathway nodes is required for the comprehensive analysis of the pathway activation profiles. We proposed an algorithm that identifies functional roles of the pathway components and applied it to annotate 3,044 human molecular pathways extracted from the Biocarta, Reactome, KEGG, Qiagen Pathway Central, NCI, and HumanCYC databases and including 9,022 gene products. The resulting knowledgebase can be applied for the direct calculation of the PALs and establishing large scale profiles of the signaling, metabolic, and DNA repair pathway regulation using high throughput gene expression data. We also provide a bioinformatic tool for PAL data calculations using the current pathway knowledgebase.
Keywords: DNA repair pathways; functional algorithmic annotation; human molecular pathway regulation; metabolic pathways; proteomics; signaling pathways; transcriptomics.
Copyright © 2021 Sorokin, Borisov, Kuzmin, Gudkov, Zolotovskaia, Garazha and Buzdin.
Conflict of interest statement
MS, AGa, and AB have a financial relationship with OmicsWay Corp.The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.The handling editor declared a shared affiliation with the authors AB, MS, and AGu at the time of review.
Figures

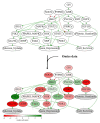
Similar articles
-
OncoboxPD: human 51 672 molecular pathways database with tools for activity calculating and visualization.Comput Struct Biotechnol J. 2022 May 10;20:2280-2291. doi: 10.1016/j.csbj.2022.05.006. eCollection 2022. Comput Struct Biotechnol J. 2022. PMID: 35615022 Free PMC article.
-
De novo assembly and functional annotation of Myrciaria dubia fruit transcriptome reveals multiple metabolic pathways for L-ascorbic acid biosynthesis.BMC Genomics. 2015 Nov 24;16:997. doi: 10.1186/s12864-015-2225-6. BMC Genomics. 2015. PMID: 26602763 Free PMC article.
-
Using proteomic and transcriptomic data to assess activation of intracellular molecular pathways.Adv Protein Chem Struct Biol. 2021;127:1-53. doi: 10.1016/bs.apcsb.2021.02.005. Epub 2021 Apr 5. Adv Protein Chem Struct Biol. 2021. PMID: 34340765 Review.
-
The reactome pathway knowledgebase.Nucleic Acids Res. 2020 Jan 8;48(D1):D498-D503. doi: 10.1093/nar/gkz1031. Nucleic Acids Res. 2020. PMID: 31691815 Free PMC article.
-
ESEA: Discovering the Dysregulated Pathways based on Edge Set Enrichment Analysis.Sci Rep. 2015 Aug 12;5:13044. doi: 10.1038/srep13044. Sci Rep. 2015. PMID: 26267116 Free PMC article.
Cited by
-
M6A "Writer" Gene METTL14: A Favorable Prognostic Biomarker and Correlated With Immune Infiltrates in Rectal Cancer.Front Oncol. 2021 Jun 17;11:615296. doi: 10.3389/fonc.2021.615296. eCollection 2021. Front Oncol. 2021. PMID: 34221955 Free PMC article.
-
Lapatinib-induced enhancement of mitochondrial respiration in HER2-positive SK-BR-3 cells: mechanism revealed by analysis of proteomic but not transcriptomic data.Front Mol Biosci. 2024 Sep 30;11:1470496. doi: 10.3389/fmolb.2024.1470496. eCollection 2024. Front Mol Biosci. 2024. PMID: 39403185 Free PMC article.
-
Molecular programs of fibrotic change in aging human lung.Nat Commun. 2021 Nov 2;12(1):6309. doi: 10.1038/s41467-021-26603-2. Nat Commun. 2021. PMID: 34728633 Free PMC article.
-
Human Blood Serum Can Diminish EGFR-Targeted Inhibition of Squamous Carcinoma Cell Growth through Reactivation of MAPK and EGFR Pathways.Cells. 2023 Aug 8;12(16):2022. doi: 10.3390/cells12162022. Cells. 2023. PMID: 37626832 Free PMC article.
-
OncoboxPD: human 51 672 molecular pathways database with tools for activity calculating and visualization.Comput Struct Biotechnol J. 2022 May 10;20:2280-2291. doi: 10.1016/j.csbj.2022.05.006. eCollection 2022. Comput Struct Biotechnol J. 2022. PMID: 35615022 Free PMC article.
References
-
- Aliper A. M., Csoka A. B., Buzdin A., Jetka T., Roumiantsev S., Moskalev A., et al. . (2015). Signaling pathway activation drift during aging: Hutchinson-Gilford Progeria Syndrome fibroblasts are comparable to normal middle-age and old-age cells. Aging 7, 26–37. 10.18632/aging.100717, PMID: - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

