Integron activity accelerates the evolution of antibiotic resistance
- PMID: 33634790
- PMCID: PMC8024014
- DOI: 10.7554/eLife.62474
Integron activity accelerates the evolution of antibiotic resistance
Abstract
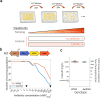
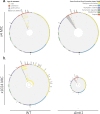
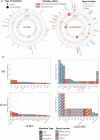
Mobile integrons are widespread genetic platforms that allow bacteria to modulate the expression of antibiotic resistance cassettes by shuffling their position from a common promoter. Antibiotic stress induces the expression of an integrase that excises and integrates cassettes, and this unique recombination and expression system is thought to allow bacteria to 'evolve on demand' in response to antibiotic pressure. To test this hypothesis, we inserted a custom three-cassette integron into Pseudomonas aeruginosa and used experimental evolution to measure the impact of integrase activity on adaptation to gentamicin. Crucially, integrase activity accelerated evolution by increasing the expression of a gentamicin resistance cassette through duplications and by eliminating redundant cassettes. Importantly, we found no evidence of deleterious off-target effects of integrase activity. In summary, integrons accelerate resistance evolution by rapidly generating combinatorial variation in cassette composition while maintaining genomic integrity.
Keywords: P. aeruginosa; antibiotic resistance; evolutionary biology; experimental evolution; infectious disease; integron; microbiology.
Plain language summary
From urinary tract infections to bacterial pneumonia, many diseases can now be treated through a course of antibiotics. Yet bacteria have evolved to respond to this threat, gaining new antibiotic resistance genes that allow them to evade the drugs. Addressing this growing issue requires to either discover new antibiotics, or to stop resistance before it emerges – a strategy that can only work if scientists know exactly how this mechanism takes place. For bacteria, it is a waste of resources to produce the proteins that confer resistance if antibiotics are absent. In fact, doing so can decrease their chance to survive and reproduce. A genetic element known as an integron can help to manage that burden. This piece of genetic information is formed of a succession of ‘cassettes’ containing antibiotic resistance genes. More proteins are made from the genes present at the start of the integron, compared to the ones towards the end. When bacteria encounter antibiotics, an enzyme called integrase is activated, allowing the organisms to shuffle the order of their cassettes in the integron. It is thought – but not yet proven – that this mechanism helps bacteria to activate their resistance ‘on demand’. To find out, Souque et al. engineered the bacteria Pseudomonas aeruginosa to carry a custom integron with three cassettes, each helping the organism to resist to a different antibiotic. In addition, only half of the bacteria had a working integrase and could therefore shuffle their gene cassettes. The organisms were then exposed to an increasing amount of the antibiotics for which the cassette in the last position provided resistance. The bacteria with a working integrase survived longer than those without, as they were able to shuffle their cassettes and move the useful antibiotic resistance gene into top position. In addition, the cassettes carrying the genes to resist to other types of antibiotics were excised from the genetic information and lost. Understanding integrons could guide future antibiotic treatment strategies, for instance by combining antibiotics with chemicals that block integrase activity. It might also be possible to force bacteria to delete resistance cassettes by cycling through different antibiotics.
© 2021, Souque et al.
Conflict of interest statement
CS, JE, RM No competing interests declared
Figures




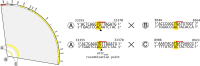



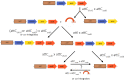
Comment in
-
Bacterial evolution on demand.Elife. 2021 Apr 6;10:e68070. doi: 10.7554/eLife.68070. Elife. 2021. PMID: 33820602 Free PMC article.
Similar articles
-
Integrons: the hidden architects of bacterial adaptation, evolution, and the challenges of antimicrobial resistance.Antonie Van Leeuwenhoek. 2025 Jun 12;118(7):90. doi: 10.1007/s10482-025-02103-x. Antonie Van Leeuwenhoek. 2025. PMID: 40506592 Review.
-
Off-Target Integron Activity Leads to Rapid Plasmid Compensatory Evolution in Response to Antibiotic Selection Pressure.mBio. 2023 Apr 25;14(2):e0253722. doi: 10.1128/mbio.02537-22. Epub 2023 Feb 22. mBio. 2023. PMID: 36840554 Free PMC article.
-
Diversity of Class 1 Integron Gene Cassette Rearrangements Selected under Antibiotic Pressure.J Bacteriol. 2015 Jul;197(13):2171-2178. doi: 10.1128/JB.02455-14. Epub 2015 Apr 20. J Bacteriol. 2015. PMID: 25897031 Free PMC article.
-
The Stringent Response Promotes Antibiotic Resistance Dissemination by Regulating Integron Integrase Expression in Biofilms.mBio. 2016 Aug 16;7(4):e00868-16. doi: 10.1128/mBio.00868-16. mBio. 2016. PMID: 27531906 Free PMC article.
-
Resistance integrons and super-integrons.Clin Microbiol Infect. 2004 Apr;10(4):272-88. doi: 10.1111/j.1198-743X.2004.00858.x. Clin Microbiol Infect. 2004. PMID: 15059115 Review.
Cited by
-
Integrons: the hidden architects of bacterial adaptation, evolution, and the challenges of antimicrobial resistance.Antonie Van Leeuwenhoek. 2025 Jun 12;118(7):90. doi: 10.1007/s10482-025-02103-x. Antonie Van Leeuwenhoek. 2025. PMID: 40506592 Review.
-
Global genomic epidemiology of bla GES-5 carbapenemase-associated integrons.Microb Genom. 2024 Dec;10(12):001312. doi: 10.1099/mgen.0.001312. Microb Genom. 2024. PMID: 39630499 Free PMC article.
-
Beav: a bacterial genome and mobile element annotation pipeline.mSphere. 2024 Aug 28;9(8):e0020924. doi: 10.1128/msphere.00209-24. Epub 2024 Jul 22. mSphere. 2024. PMID: 39037262 Free PMC article.
-
Within-patient evolution of plasmid-mediated antimicrobial resistance.Nat Ecol Evol. 2022 Dec;6(12):1980-1991. doi: 10.1038/s41559-022-01908-7. Epub 2022 Oct 27. Nat Ecol Evol. 2022. PMID: 36303001 Free PMC article.
-
Detection of IMP-4 and SFO-1 co-producing ST51 Enterobacter hormaechei clinical isolates.Front Cell Infect Microbiol. 2022 Oct 27;12:998578. doi: 10.3389/fcimb.2022.998578. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 36389152 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

