Epigenomic characterization of latent HIV infection identifies latency regulating transcription factors
- PMID: 33635929
- PMCID: PMC7946360
- DOI: 10.1371/journal.ppat.1009346
Epigenomic characterization of latent HIV infection identifies latency regulating transcription factors
Abstract
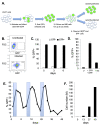
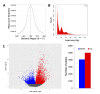
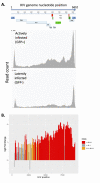
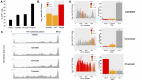
Transcriptional silencing of HIV in CD4 T cells generates a reservoir of latently infected cells that can reseed infection after interruption of therapy. As such, these cells represent the principal barrier to curing HIV infection, but little is known about their characteristics. To further our understanding of the molecular mechanisms of latency, we characterized a primary cell model of HIV latency in which infected cells adopt heterogeneous transcriptional fates. In this model, we observed that latency is a stable, heritable state that is transmitted through cell division. Using Assay of Transposon-Accessible Chromatin sequencing (ATACseq) we found that latently infected cells exhibit greatly reduced proviral accessibility, indicating the presence of chromatin-based structural barriers to viral gene expression. By quantifying the activity of host cell transcription factors, we observe elevated activity of Forkhead and Kruppel-like factor transcription factors (TFs), and reduced activity of AP-1, RUNX and GATA TFs in latently infected cells. Interestingly, latency reversing agents with different mechanisms of action caused distinct patterns of chromatin reopening across the provirus. We observe that binding sites for the chromatin insulator CTCF are highly enriched in the differentially open chromatin of infected CD4 T cells. Furthermore, depletion of CTCF inhibited HIV latency, identifying this factor as playing a key role in the initiation or enforcement of latency. These data indicate that HIV latency develops preferentially in cells with a distinct pattern of TF activity that promotes a closed proviral structure and inhibits viral gene expression. Furthermore, these findings identify CTCF as a novel regulator of HIV latency.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

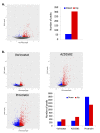


References
-
- Serrano-Villar S, Sainz T, Lee SA, Hunt PW, Sinclair E, Shacklett BL, et al. HIV-Infected Individuals with Low CD4/CD8 Ratio despite Effective Antiretroviral Therapy Exhibit Altered T Cell Subsets, Heightened CD8+ T Cell Activation, and Increased Risk of Non-AIDS Morbidity and Mortality. PLoS Pathog. 2014;10. 10.1371/journal.ppat.1004078 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

