Uncovering candidate genes involved in photosynthetic capacity using unexplored genetic variation in Spring Wheat
- PMID: 33638599
- PMCID: PMC8384606
- DOI: 10.1111/pbi.13568
Uncovering candidate genes involved in photosynthetic capacity using unexplored genetic variation in Spring Wheat
Abstract
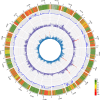
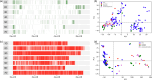
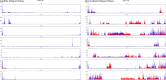
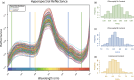
To feed an ever-increasing population we must leverage advances in genomics and phenotyping to harness the variation in wheat breeding populations for traits like photosynthetic capacity which remains unoptimized. Here we survey a diverse set of wheat germplasm containing elite, introgression and synthetic derivative lines uncovering previously uncharacterized variation. We demonstrate how strategic integration of exotic material alleviates the D genome genetic bottleneck in wheat, increasing SNP rate by 62% largely due to Ae. tauschii synthetic wheat donors. Across the panel, 67% of the Ae. tauschii donor genome is represented as introgressions in elite backgrounds. We show how observed genetic variation together with hyperspectral reflectance data can be used to identify candidate genes for traits relating to photosynthetic capacity using association analysis. This demonstrates the value of genomic methods in uncovering hidden variation in wheat and how that variation can assist breeding efforts and increase our understanding of complex traits.
Keywords: Aegilops Tauschii; Triticum aestivum; GWAS; capture sequencing; exotic material; hyperspectral reflectance.
© 2021 The Authors. Plant Biotechnology Journal published by Society for Experimental Biology and The Association of Applied Biologists and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
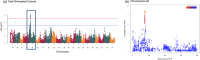
Similar articles
-
A population of wheat multiple synthetic derivatives: an effective platform to explore, harness and utilize genetic diversity of Aegilops tauschii for wheat improvement.Theor Appl Genet. 2018 Aug;131(8):1615-1626. doi: 10.1007/s00122-018-3102-x. Epub 2018 Apr 28. Theor Appl Genet. 2018. PMID: 29705916 Free PMC article.
-
Unlocking the novel genetic diversity and population structure of synthetic Hexaploid wheat.BMC Genomics. 2018 Aug 6;19(1):591. doi: 10.1186/s12864-018-4969-2. BMC Genomics. 2018. PMID: 30081829 Free PMC article.
-
Development of a set of Triticum aestivum-Aegilops tauschii introgression lines.Hereditas. 2001;135(2-3):139-43. doi: 10.1111/j.1601-5223.2001.00139.x. Hereditas. 2001. PMID: 12152326
-
Hyperspectral reflectance-based phenotyping for quantitative genetics in crops: Progress and challenges.Plant Commun. 2021 May 27;2(4):100209. doi: 10.1016/j.xplc.2021.100209. eCollection 2021 Jul 12. Plant Commun. 2021. PMID: 34327323 Free PMC article. Review.
-
The Resurgence of Introgression Breeding, as Exemplified in Wheat Improvement.Front Plant Sci. 2020 Mar 6;11:252. doi: 10.3389/fpls.2020.00252. eCollection 2020. Front Plant Sci. 2020. PMID: 32211007 Free PMC article. Review.
Cited by
-
Effect of Flowering Time-Related Genes on Biomass, Harvest Index, and Grain Yield in CIMMYT Elite Spring Bread Wheat.Biology (Basel). 2021 Sep 1;10(9):855. doi: 10.3390/biology10090855. Biology (Basel). 2021. PMID: 34571732 Free PMC article.
-
Water stress effects on stay green and chlorophyll fluorescence with focus on yield characteristics of diverse bread wheats.Planta. 2023 Apr 28;257(6):104. doi: 10.1007/s00425-023-04140-0. Planta. 2023. PMID: 37115268
-
A model-guided holistic review of exploiting natural variation of photosynthesis traits in crop improvement.J Exp Bot. 2022 May 23;73(10):3173-3188. doi: 10.1093/jxb/erac109. J Exp Bot. 2022. PMID: 35323898 Free PMC article. Review.
-
Improving Photosynthetic Metabolism for Crop Yields: What Is Going to Work?Front Plant Sci. 2021 Sep 21;12:743862. doi: 10.3389/fpls.2021.743862. eCollection 2021. Front Plant Sci. 2021. PMID: 34621287 Free PMC article. No abstract available.
-
Exotic alleles contribute to heat tolerance in wheat under field conditions.Commun Biol. 2023 Jan 9;6(1):21. doi: 10.1038/s42003-022-04325-5. Commun Biol. 2023. PMID: 36624201 Free PMC article.
References
-
- Allen, A.M. , Winfield, M.O. , Burridge, A.J. , Downie, R.C. , Benbow, H.R. , Barker, G.L.A. , Wilkinson, P.A. et al. (2016) Characterization of a Wheat Breeders’ Array suitable for high‐throughput SNP genotyping of global accessions of hexaploid bread wheat (Triticum aestivum). Plant Biotechnol. J. 15, 390–401. - PMC - PubMed
-
- Alvarado, G. , López, M. , Vargas, M. , Pacheco, Á. , Rodríguez, F. , Burgueño, J. and Crossa, J. (2019). META‐R (Multi Environment Trail Analysis with R for Windows) Version 6.04.
-
- Araus, J.‐L. and Cairns, J.E. (2014) Field high‐throughput phenotyping: the new crop breeding frontier. Trends Plant Sci. 19, 52–61. - PubMed
-
- Bhusal, N. , Sharma, P. , Sareen, S. and Sarial, A.K. (2018) Mapping QTLs for chlorophyll content and chlorophyll fluorescence in wheat under heat stress. Biol. Plant. 62, 721–731.
-
- Blackburn, G.A. (2006) Hyperspectral remote sensing of plant pigments. J. Exp. Bot. 58, 855–867. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

