Multifaceted Mechanism of Amicoumacin A Inhibition of Bacterial Translation
- PMID: 33643246
- PMCID: PMC7907450
- DOI: 10.3389/fmicb.2021.618857
Multifaceted Mechanism of Amicoumacin A Inhibition of Bacterial Translation
Abstract
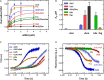
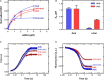
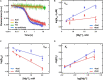
Amicoumacin A (Ami) halts bacterial growth by inhibiting the ribosome during translation. The Ami binding site locates in the vicinity of the E-site codon of mRNA. However, Ami does not clash with mRNA, rather stabilizes it, which is relatively unusual and implies a unique way of translation inhibition. In this work, we performed a kinetic and thermodynamic investigation of Ami influence on the main steps of polypeptide synthesis. We show that Ami reduces the rate of the functional canonical 70S initiation complex (IC) formation by 30-fold. Additionally, our results indicate that Ami promotes the formation of erroneous 30S ICs; however, IF3 prevents them from progressing towards translation initiation. During early elongation steps, Ami does not compromise EF-Tu-dependent A-site binding or peptide bond formation. On the other hand, Ami reduces the rate of peptidyl-tRNA movement from the A to the P site and significantly decreases the amount of the ribosomes capable of polypeptide synthesis. Our data indicate that Ami progressively decreases the activity of translating ribosomes that may appear to be the main inhibitory mechanism of Ami. Indeed, the use of EF-G mutants that confer resistance to Ami (G542V, G581A, or ins544V) leads to a complete restoration of the ribosome functionality. It is possible that the changes in translocation induced by EF-G mutants compensate for the activity loss caused by Ami.
Keywords: amicoumacin A; antibiotic resistance; elongation factor EF-G; initiation; microscale thermophoresis; rapid kinetics; translocation.
Copyright © 2021 Maksimova, Vinogradova, Osterman, Kasatsky, Nikonov, Milón, Dontsova, Sergiev, Paleskava and Konevega.
Conflict of interest statement
AK is a founder of the company NanoTemper Technologies Rus (St. Petersburg, Russia), which provides services and devices based on MST and nanoDSF and represents NanoTemper Technologies GmbH (Germany). The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
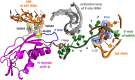
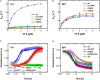

References
-
- Azumi M., Ogawa K., Fujita T., Takeshita M., Yoshida R., Furumai T., et al. (2008). Bacilosarcins A and B, novel bioactive isocoumarins with unusual heterocyclic cores from the marine-derived bacterium Bacillus subtilis. Tetrahedron 64 6420–6425. 10.1016/j.tet.2008.04.076 - DOI
-
- Dinos G., Wilson D. N., Teraoka Y., Szaflarski W., Fucini P., Kalpaxis D., et al. (2004). Dissecting the ribosomal inhibition mechanisms of edeine and pactamycin: the universally conserved residues G693 and C795 regulate P-site RNA binding. Mol. Cell. 13 113–124. 10.1016/s1097-2765(04)00002-4 - DOI - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

