The relationship between donor-recipient genetic distance and long-term kidney transplant outcome
- PMID: 33655195
- PMCID: PMC7888353
- DOI: 10.12688/hrbopenres.13021.1
The relationship between donor-recipient genetic distance and long-term kidney transplant outcome
Abstract
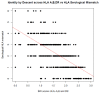
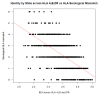
Background: We set out to quantify shared genetic ancestry between unrelated kidney donor-recipient pairs and test it as a predictor of time to graft failure. Methods: In a homogenous, unrelated, European cohort of deceased-donor kidney transplant pairs (n pairs = 1,808), we calculated, using common genetic variation, shared ancestry at the genic (n loci=40,053) and genomic level. We conducted a sub-analysis focused on transmembrane protein coding genes (n transcripts=8,637) and attempted replication of a previously published nonsynonymous transmembrane mismatch score. Measures of shared genetic ancestry were tested in a survival model against time to death-censored graft failure. Results: Shared ancestry calculated across the human leukocyte antigen (HLA) significantly associated with graft survival in individuals who had a high serological mismatch (n pairs = 186) with those who did not have any HLA mismatches indicating that shared ancestry calculated specific loci can capture known associations with genes impacting graft outcome. None of the other measures of shared ancestry at a genic level, genome-wide scale, transmembrane subset or nonsynonymous transmembrane mismatch score analysis were significant predictors of time to graft failure. Conclusions: In a large unrelated, deceased-donor European ancestry renal transplant cohort, shared donor-recipient genetic ancestry, calculated using common genetic variation, has limited value in predicting transplant outcome both on a genomic scale and at a genic level (other than at the HLA loci).
Keywords: HLA; graft failure; identity-by-descent; identity-by-state; kidney transplant; shared genetic ancestry; transplant.
Copyright: © 2020 Stapleton CP et al.
Conflict of interest statement
No competing interests were disclosed.
Figures
Similar articles
-
Donor-Recipient Non-HLA Variants, Mismatches and Renal Allograft Outcomes: Evolving Paradigms.Front Immunol. 2022 Apr 1;13:822353. doi: 10.3389/fimmu.2022.822353. eCollection 2022. Front Immunol. 2022. PMID: 35432337 Free PMC article. Review.
-
Effect of amino acid mismatch in the UNOS dataset.Clin Transpl. 2011:299-309. Clin Transpl. 2011. PMID: 22755423
-
Genome-wide non-HLA donor-recipient genetic differences influence renal allograft survival via early allograft fibrosis.Kidney Int. 2020 Sep;98(3):758-768. doi: 10.1016/j.kint.2020.04.039. Epub 2020 May 23. Kidney Int. 2020. PMID: 32454123 Free PMC article.
-
Contribution of non-HLA incompatibility between donor and recipient to kidney allograft survival: genome-wide analysis in a prospective cohort.Lancet. 2019 Mar 2;393(10174):910-917. doi: 10.1016/S0140-6736(18)32473-5. Epub 2019 Feb 14. Lancet. 2019. PMID: 30773281
-
Risk Factors for 1-Year Graft Loss After Kidney Transplantation: Systematic Review and Meta-Analysis.Clin J Am Soc Nephrol. 2019 Nov 7;14(11):1642-1650. doi: 10.2215/CJN.05560519. Epub 2019 Sep 20. Clin J Am Soc Nephrol. 2019. PMID: 31540931 Free PMC article.
Cited by
-
Donor-Recipient Non-HLA Variants, Mismatches and Renal Allograft Outcomes: Evolving Paradigms.Front Immunol. 2022 Apr 1;13:822353. doi: 10.3389/fimmu.2022.822353. eCollection 2022. Front Immunol. 2022. PMID: 35432337 Free PMC article. Review.
-
Donor and Recipient Polygenic Risk Scores Influence Kidney Transplant Function.Transpl Int. 2025 Mar 4;38:14171. doi: 10.3389/ti.2025.14171. eCollection 2025. Transpl Int. 2025. PMID: 40104404 Free PMC article.
-
Improving long-term kidney allograft survival by rethinking HLA compatibility: from molecular matching to non-HLA genes.Front Genet. 2024 Oct 2;15:1442018. doi: 10.3389/fgene.2024.1442018. eCollection 2024. Front Genet. 2024. PMID: 39415982 Free PMC article. Review.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

