Aberrant Gcm1 expression mediates Wnt/β-catenin pathway activation in folate deficiency involved in neural tube defects
- PMID: 33664222
- PMCID: PMC7933360
- DOI: 10.1038/s41419-020-03313-z
Aberrant Gcm1 expression mediates Wnt/β-catenin pathway activation in folate deficiency involved in neural tube defects
Abstract
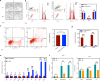
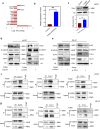
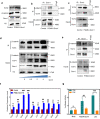
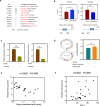
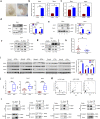
Wnt signaling plays a major role in early neural development. An aberrant activation in Wnt/β-catenin pathway causes defective anteroposterior patterning, which results in neural tube closure defects (NTDs). Changes in folate metabolism may participate in early embryo fate determination. We have identified that folate deficiency activated Wnt/β-catenin pathway by upregulating a chorion-specific transcription factor Gcm1. Specifically, folate deficiency promoted formation of the Gcm1/β-catenin/T-cell factor (TCF4) complex formation to regulate the Wnt targeted gene transactivation through Wnt-responsive elements. Moreover, the transcription factor Nanog upregulated Gcm1 transcription in mESCs under folate deficiency. Lastly, in NTDs mouse models and low-folate NTDs human brain samples, Gcm1 and Wnt/β-catenin targeted genes related to neural tube closure are specifically overexpressed. These results indicated that low-folate level promoted Wnt/β-catenin signaling via activating Gcm1, and thus leaded into aberrant vertebrate neural development.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures

Similar articles
-
β-catenin regulates Pax3 and Cdx2 for caudal neural tube closure and elongation.Development. 2014 Jan;141(1):148-57. doi: 10.1242/dev.101550. Epub 2013 Nov 27. Development. 2014. PMID: 24284205 Free PMC article.
-
High levels of iron supplementation prevents neural tube defects in the Fpn1ffe mouse model.Birth Defects Res. 2017 Jan 30;109(2):81-91. doi: 10.1002/bdra.23542. Birth Defects Res. 2017. PMID: 28008752 Free PMC article.
-
Inositol, neural tube closure and the prevention of neural tube defects.Birth Defects Res. 2017 Jan 30;109(2):68-80. doi: 10.1002/bdra.23533. Birth Defects Res. 2017. PMID: 27324558 Free PMC article. Review.
-
The effect of folic acid deficiency on Mest/Peg1 in neural tube defects.Int J Neurosci. 2021 May;131(5):468-477. doi: 10.1080/00207454.2020.1750386. Epub 2020 Apr 16. Int J Neurosci. 2021. PMID: 32241207
-
Perspectives on folate with special reference to epigenetics and neural tube defects.Reprod Toxicol. 2024 Apr;125:108576. doi: 10.1016/j.reprotox.2024.108576. Epub 2024 Mar 12. Reprod Toxicol. 2024. PMID: 38479591 Review.
Cited by
-
METTL3 shapes m6A epitranscriptomic landscape for successful human placentation.bioRxiv [Preprint]. 2024 Jul 13:2024.07.12.603294. doi: 10.1101/2024.07.12.603294. bioRxiv. 2024. PMID: 39026770 Free PMC article. Preprint.
-
Melatonin alleviates valproic acid-induced neural tube defects by modulating Src/PI3K/ERK signaling and oxidative stress.Acta Biochim Biophys Sin (Shanghai). 2024 Jan 25;56(1):23-33. doi: 10.3724/abbs.2023234. Acta Biochim Biophys Sin (Shanghai). 2024. PMID: 38062774 Free PMC article.
-
Spatiotemporal protein dynamics during early organogenesis in mouse conceptuses treated with valproic acid.Neurotoxicol Teratol. 2023 Sep-Oct;99:107286. doi: 10.1016/j.ntt.2023.107286. Epub 2023 Jul 11. Neurotoxicol Teratol. 2023. PMID: 37442398 Free PMC article.
-
A quest for genetic causes underlying signaling pathways associated with neural tube defects.Front Pediatr. 2023 May 22;11:1126209. doi: 10.3389/fped.2023.1126209. eCollection 2023. Front Pediatr. 2023. PMID: 37284286 Free PMC article. Review.
-
Folic acid alleviates the negative effects of dexamethasone induced stress on production performance in Hyline Brown laying hens.Anim Nutr. 2024 Dec 14;20:54-65. doi: 10.1016/j.aninu.2024.11.011. eCollection 2025 Mar. Anim Nutr. 2024. PMID: 39949729 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

