Prediction of neo-epitope immunogenicity reveals TCR recognition determinants and provides insight into immunoediting
- PMID: 33665637
- PMCID: PMC7897774
- DOI: 10.1016/j.xcrm.2021.100194
Prediction of neo-epitope immunogenicity reveals TCR recognition determinants and provides insight into immunoediting
Abstract
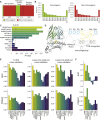
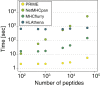
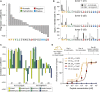
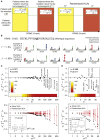
CD8+ T cell recognition of peptide epitopes plays a central role in immune responses against pathogens and tumors. However, the rules that govern which peptides are truly recognized by existing T cell receptors (TCRs) remain poorly understood, precluding accurate predictions of neo-epitopes for cancer immunotherapy. Here, we capitalize on recent (neo-)epitope data to train a predictor of immunogenic epitopes (PRIME), which captures molecular properties of both antigen presentation and TCR recognition. PRIME not only improves prioritization of neo-epitopes but also correlates with T cell potency and unravels biophysical determinants of TCR recognition that we experimentally validate. Analysis of cancer genomics data reveals that recurrent mutations tend to be less frequent in patients where they are predicted to be immunogenic, providing further evidence for immunoediting in human cancer. PRIME will facilitate identification of pathogen epitopes in infectious diseases and neo-epitopes in cancer immunotherapy.
Keywords: TCR recognition; immunoediting; immunogenicity; neo-epitope predictions; tumor immunology.
© 2021 The Author(s).
Conflict of interest statement
The authors declare no competing interests. G. Ciriello is a member of the advisory board of Cell Reports Medicine.
Figures



References
-
- Sahin U., Derhovanessian E., Miller M., Kloke B.-P., Simon P., Löwer M., Bukur V., Tadmor A.D., Luxemburger U., Schrörs B. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature. 2017;547:222–226. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

