Underground metabolism facilitates the evolution of novel pathways for vitamin B6 biosynthesis
- PMID: 33665688
- PMCID: PMC7954711
- DOI: 10.1007/s00253-021-11199-w
Underground metabolism facilitates the evolution of novel pathways for vitamin B6 biosynthesis
Abstract
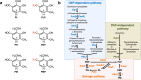
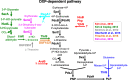
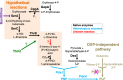
The term vitamin B6 is a designation for the vitamers pyridoxal, pyridoxamine, pyridoxine and the respective phosphate esters pyridoxal-5'-phosphate (PLP), pyridoxamine-5'-phosphate and pyridoxine-5'-phosphate. Animals and humans are unable to synthesise vitamin B6. These organisms have to take up vitamin B6 with their diet. Therefore, vitamin B6 is of commercial interest as a food additive and for applications in the pharmaceutical industry. As yet, two naturally occurring routes for de novo synthesis of PLP are known. Both routes have been genetically engineered to obtain bacteria overproducing vitamin B6. Still, major genetic engineering efforts using the existing pathways are required for developing fermentation processes that could outcompete the chemical synthesis of vitamin B6. Recent suppressor screens using mutants of the Gram-negative and Gram-positive model bacteria Escherichia coli and Bacillus subtilis, respectively, carrying mutations in the native pathways or heterologous genes uncovered novel routes for PLP biosynthesis. These pathways consist of promiscuous enzymes and enzymes that are already involved in vitamin B6 biosynthesis. Thus, E. coli and B. subtilis contain multiple promiscuous enzymes causing a so-called underground metabolism allowing the bacteria to bypass disrupted vitamin B6 biosynthetic pathways. The suppressor screens also show the genomic plasticity of the bacteria to suppress a genetic lesion. We discuss the potential of the serendipitous pathways to serve as a starting point for the development of bacteria overproducing vitamin B6. KEY POINTS: • Known vitamin B6 routes have been genetically engineered. • Underground metabolism facilitates the emergence of novel vitamin B6 biosynthetic pathways. • These pathways may be suitable to engineer bacteria overproducing vitamin B6.
Keywords: Enzyme promiscuity; Enzyme specificity; Genetic suppression; Promiscuous enzyme; Pyridoxal 5′-phosphate.
Conflict of interest statement
The authors declare no competing interests.
Figures
Similar articles
-
Vitamin B6 metabolism in microbes and approaches for fermentative production.Biotechnol Adv. 2017 Jan-Feb;35(1):31-40. doi: 10.1016/j.biotechadv.2016.11.004. Epub 2016 Nov 24. Biotechnol Adv. 2017. PMID: 27890703 Review.
-
A two-step evolutionary process establishes a non-native vitamin B6 pathway in Bacillus subtilis.Environ Microbiol. 2018 Jan;20(1):156-168. doi: 10.1111/1462-2920.13950. Epub 2017 Oct 27. Environ Microbiol. 2018. PMID: 29027347
-
The Role of YggS in Vitamin B6 Homeostasis in Salmonella enterica Is Informed by Heterologous Expression of Yeast SNZ3.J Bacteriol. 2020 Oct 22;202(22):e00383-20. doi: 10.1128/JB.00383-20. Print 2020 Oct 22. J Bacteriol. 2020. PMID: 32900833 Free PMC article.
-
Mechanism of Pyridoxine 5'-Phosphate Accumulation in Pyridoxal 5'-Phosphate-Binding Protein Deficiency.J Bacteriol. 2022 Mar 15;204(3):e0052121. doi: 10.1128/JB.00521-21. Epub 2022 Jan 3. J Bacteriol. 2022. PMID: 34978460 Free PMC article.
-
Phylogenetic analyses and comparative genomics of vitamin B6 (pyridoxine) and pyridoxal phosphate biosynthesis pathways.J Mol Microbiol Biotechnol. 2001 Jan;3(1):1-20. J Mol Microbiol Biotechnol. 2001. PMID: 11200221 Review.
Cited by
-
The mosquito Aedes aegypti requires a gut microbiota for normal fecundity, longevity and vector competence.Commun Biol. 2023 Nov 13;6(1):1154. doi: 10.1038/s42003-023-05545-z. Commun Biol. 2023. PMID: 37957247 Free PMC article.
-
Protein engineering and iterative multimodule optimization for vitamin B6 production in Escherichia coli.Nat Commun. 2023 Aug 31;14(1):5304. doi: 10.1038/s41467-023-40928-0. Nat Commun. 2023. PMID: 37652926 Free PMC article.
-
Vitamins as regulators of calcium-containing kidney stones - new perspectives on the role of the gut microbiome.Nat Rev Urol. 2023 Oct;20(10):615-637. doi: 10.1038/s41585-023-00768-5. Epub 2023 May 9. Nat Rev Urol. 2023. PMID: 37161031 Free PMC article. Review.
-
Establishment of age-stratified serum vitamin B6 pyridoxal and pyridoxic acid reference intervals among children aged 1-16 years in Henan, china.Front Nutr. 2025 Jul 30;9:1592045. doi: 10.3389/fnut.2025.1592045. eCollection 2025. Front Nutr. 2025. PMID: 40810010 Free PMC article.
-
Cronobacter sakazakii Pyridoxal Kinase PdxY Mediated by TreR and pESA3 Is Essential for Vitamin B6 (PLP) Maintenance and Virulence.Appl Environ Microbiol. 2023 Aug 30;89(8):e0092423. doi: 10.1128/aem.00924-23. Epub 2023 Jul 17. Appl Environ Microbiol. 2023. PMID: 37458600 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

