Interactions of Sea Anemone Toxins with Insect Sodium Channel-Insights from Electrophysiology and Molecular Docking Studies
- PMID: 33670972
- PMCID: PMC7957711
- DOI: 10.3390/molecules26051302
Interactions of Sea Anemone Toxins with Insect Sodium Channel-Insights from Electrophysiology and Molecular Docking Studies
Abstract
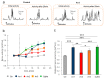
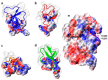
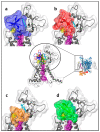
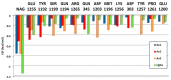
Animal venoms are considered as a promising source of new drugs. Sea anemones release polypeptides that affect electrical activity of neurons of their prey. Voltage dependent sodium (Nav) channels are the common targets of Av1, Av2, and Av3 toxins from Anemonia viridis and CgNa from Condylactis gigantea. The toxins bind to the extracellular side of a channel and slow its fast inactivation, but molecular details of the binding modes are not known. Electrophysiological measurements on Periplaneta americana neuronal preparation revealed differences in potency of these toxins to increase nerve activity. Av1 and CgNa exhibit the strongest effects, while Av2 the weakest effect. Extensive molecular docking using a modern SMINA computer method revealed only partial overlap among the sets of toxins' and channel's amino acid residues responsible for the selectivity and binding modes. Docking positions support earlier supposition that the higher neuronal activity observed in electrophysiology should be attributed to hampering the fast inactivation gate by interactions of an anemone toxin with the voltage driven S4 helix from domain IV of cockroach Nav channel (NavPaS). Our modelling provides new data linking activity of toxins with their mode of binding in site 3 of NavPaS channel.
Keywords: anemone toxins; docking; electrophysiology; fast inactivation; sodium channels.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures


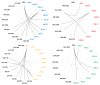

References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

