G-Quadruplex Structures Colocalize with Transcription Factories and Nuclear Speckles Surrounded by Acetylated and Dimethylated Histones H3
- PMID: 33671470
- PMCID: PMC7922289
- DOI: 10.3390/ijms22041995
G-Quadruplex Structures Colocalize with Transcription Factories and Nuclear Speckles Surrounded by Acetylated and Dimethylated Histones H3
Abstract
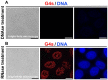
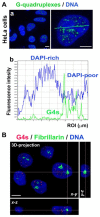
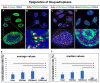
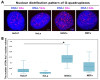
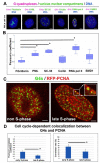
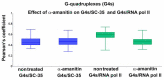
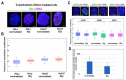
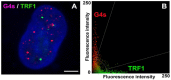
G-quadruplexes (G4s) are four-stranded helical structures that regulate several nuclear processes, including gene expression and telomere maintenance. We observed that G4s are located in GC-rich (euchromatin) regions and outside the fibrillarin-positive compartment of nucleoli. Genomic regions around G4s were preferentially H3K9 acetylated and H3K9 dimethylated, but H3K9me3 rarely decorated G4 structures. We additionally observed the variability in the number of G4s in selected human and mouse cell lines. We found the highest number of G4s in human embryonic stem cells. We observed the highest degree of colocalization between G4s and transcription factories, positive on the phosphorylated form of RNA polymerase II (RNAP II). Similarly, a high colocalization rate was between G4s and nuclear speckles, enriched in pre-mRNA splicing factor SC-35. PML bodies, the replication protein SMD1, and Cajal bodies colocalized with G4s to a lesser extent. Thus, G4 structures seem to appear mainly in nuclear compartments transcribed via RNAP II, and pre-mRNA is spliced via the SC-35 protein. However, α-amanitin, an inhibitor of RNAP II, did not affect colocalization between G4s and transcription factories as well as G4s and SC-35-positive domains. In addition, irradiation by γ-rays did not change a mutual link between G4s and DNA repair proteins (G4s/γH2AX, G4s/53BP1, and G4s/MDC1), accumulated into DNA damage foci. Described characteristics of G4s seem to be the manifestation of pronounced G4s stability that is likely maintained not only via a high-order organization of these structures but also by a specific histone signature, including H3K9me2, responsible for chromatin compaction.
Keywords: DNA repair; G-quadruplex structure; epigenetics; nuclear bodies; nuclear speckles; transcription factories.
Conflict of interest statement
The authors have declared that there are no conflict of interest.
Figures
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

