Generation and Breeding of EGFP-Transgenic Marmoset Monkeys: Cell Chimerism and Implications for Disease Modeling
- PMID: 33673402
- PMCID: PMC7996964
- DOI: 10.3390/cells10030505
Generation and Breeding of EGFP-Transgenic Marmoset Monkeys: Cell Chimerism and Implications for Disease Modeling
Abstract
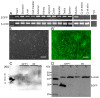
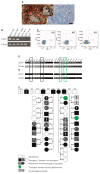
Genetic modification of non-human primates (NHP) paves the way for realistic disease models. The common marmoset is a NHP species increasingly used in biomedical research. Despite the invention of RNA-guided nucleases, one strategy for protein overexpression in NHP is still lentiviral transduction. We generated three male and one female enhanced green fluorescent protein (EGFP)-transgenic founder marmosets via lentiviral transduction of natural preimplantation embryos. All founders accomplished germline transmission of the transgene by natural mating, yielding 20 transgenic offspring together (in total, 45 pups; 44% transgenic). This demonstrates that the transgenic gametes are capable of natural fertilization even when in competition with wildtype gametes. Importantly, 90% of the transgenic offspring showed transgene silencing, which is in sharp contrast to rodents, where the identical transgene facilitated robust EGFP expression. Furthermore, we consistently discovered somatic, but so far, no germ cell chimerism in mixed wildtype/transgenic litters. Somatic cell chimerism resulted in false-positive genotyping of the respective wildtype littermates. For the discrimination of transgenic from transgene-chimeric animals by polymerase chain reaction on skin samples, a chimeric cell depletion protocol was established. In summary, it is possible to establish a cohort of genetically modified marmosets by natural mating, but specific requirements including careful promoter selection are essential.
Keywords: chimerism; embryo; genetic modification; germ cell; germline transmission; hematopoietic stem cell; marmoset monkey; non-human primate; transgenesis.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

