Recent genetic connectivity and clinal variation in chimpanzees
- PMID: 33674780
- PMCID: PMC7935964
- DOI: 10.1038/s42003-021-01806-x
Recent genetic connectivity and clinal variation in chimpanzees
Abstract
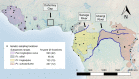
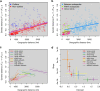
Much like humans, chimpanzees occupy diverse habitats and exhibit extensive behavioural variability. However, chimpanzees are recognized as a discontinuous species, with four subspecies separated by historical geographic barriers. Nevertheless, their range-wide degree of genetic connectivity remains poorly resolved, mainly due to sampling limitations. By analyzing a geographically comprehensive sample set amplified at microsatellite markers that inform recent population history, we found that isolation by distance explains most of the range-wide genetic structure of chimpanzees. Furthermore, we did not identify spatial discontinuities corresponding with the recognized subspecies, suggesting that some of the subspecies-delineating geographic barriers were recently permeable to gene flow. Substantial range-wide genetic connectivity is consistent with the hypothesis that behavioural flexibility is a salient driver of chimpanzee responses to changing environmental conditions. Finally, our observation of strong local differentiation associated with recent anthropogenic pressures portends future loss of critical genetic diversity if habitat fragmentation and population isolation continue unabated.
Conflict of interest statement
The authors declare no competing interests.
Figures


References
-
- Potts R. Hominin evolution in settings of strong environmental variability. Quat. Sci. Rev. 2013;73:1–13. doi: 10.1016/j.quascirev.2013.04.003. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

