Intestinal Host Response to SARS-CoV-2 Infection and COVID-19 Outcomes in Patients With Gastrointestinal Symptoms
- PMID: 33676971
- PMCID: PMC7931673
- DOI: 10.1053/j.gastro.2021.02.056
Intestinal Host Response to SARS-CoV-2 Infection and COVID-19 Outcomes in Patients With Gastrointestinal Symptoms
Abstract
Background & aims: Given that gastrointestinal (GI) symptoms are a prominent extrapulmonary manifestation of COVID-19, we investigated intestinal infection with SARS-CoV-2, its effect on pathogenesis, and clinical significance.
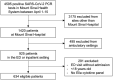
Methods: Human intestinal biopsy tissues were obtained from patients with COVID-19 (n = 19) and uninfected control individuals (n = 10) for microscopic examination, cytometry by time of flight analyses, and RNA sequencing. Additionally, disease severity and mortality were examined in patients with and without GI symptoms in 2 large, independent cohorts of hospitalized patients in the United States (N = 634) and Europe (N = 287) using multivariate logistic regressions.
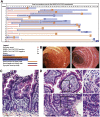
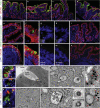
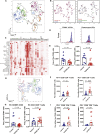
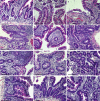
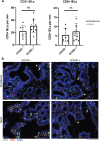
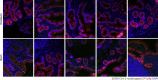
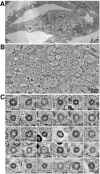
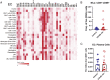
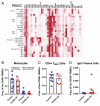
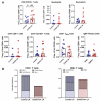
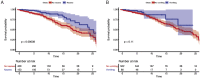
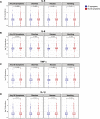
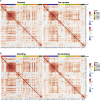
Results: COVID-19 case patients and control individuals in the biopsy cohort were comparable for age, sex, rates of hospitalization, and relevant comorbid conditions. SARS-CoV-2 was detected in small intestinal epithelial cells by immunofluorescence staining or electron microscopy in 15 of 17 patients studied. High-dimensional analyses of GI tissues showed low levels of inflammation, including down-regulation of key inflammatory genes including IFNG, CXCL8, CXCL2, and IL1B and reduced frequencies of proinflammatory dendritic cells compared with control individuals. Consistent with these findings, we found a significant reduction in disease severity and mortality in patients presenting with GI symptoms that was independent of sex, age, and comorbid illnesses and despite similar nasopharyngeal SARS-CoV-2 viral loads. Furthermore, there was reduced levels of key inflammatory proteins in circulation in patients with GI symptoms.
Conclusions: These data highlight the absence of a proinflammatory response in the GI tract despite detection of SARS-CoV-2. In parallel, reduced mortality in patients with COVID-19 presenting with GI symptoms was observed. A potential role of the GI tract in attenuating SARS-CoV-2-associated inflammation needs to be further examined.
Keywords: COVID-19; GI infection; GI symptoms; SARS-CoV-2; host immune response; outcomes.
Copyright © 2021 AGA Institute. All rights reserved.
Figures








Comment in
-
COVID-19 Gastrointestinal Symptoms and Attenuation of the Immune Response to SARS-CoV-2.Gastroenterology. 2021 Jun;160(7):2251-2254. doi: 10.1053/j.gastro.2021.03.029. Epub 2021 Mar 19. Gastroenterology. 2021. PMID: 33753106 Free PMC article. No abstract available.
References
Supplementary References
-
- Mastronarde D.N. Automated electron microscope tomography using robust prediction of specimen movements. J Struct Biol. 2005;152:36–51. - PubMed
-
- Mastronarde D.N. Correction for non-perpendicularity of beam and tilt axis in tomographic reconstructions with the IMOD package. J Microsc. 2008;230:212–217. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 DK123749/DK/NIDDK NIH HHS/United States
- U24 CA224319/CA/NCI NIH HHS/United States
- HHSN272201400008C/AI/NIAID NIH HHS/United States
- R01 AI113186/AI/NIAID NIH HHS/United States
- U01 DK124165/DK/NIDDK NIH HHS/United States
- P20 GM125498/GM/NIGMS NIH HHS/United States
- R01 CA234614/CA/NCI NIH HHS/United States
- P01 AI061093/AI/NIAID NIH HHS/United States
- F30 CA243210/CA/NCI NIH HHS/United States
- R01 DK121072/DK/NIDDK NIH HHS/United States
- R01 AI107301/AI/NIAID NIH HHS/United States
- S10 OD018522/OD/NIH HHS/United States
- S10 OD026880/OD/NIH HHS/United States
- T32 AI007605/AI/NIAID NIH HHS/United States
- P01 CA190174/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

