A Novel Familial PHP1B Variant With Incomplete Loss of Methylation at GNAS-A/B and Enhanced Methylation at GNAS-AS2
- PMID: 33677588
- PMCID: PMC8372637
- DOI: 10.1210/clinem/dgab136
A Novel Familial PHP1B Variant With Incomplete Loss of Methylation at GNAS-A/B and Enhanced Methylation at GNAS-AS2
Abstract
Context: Pseudohypoparathyroidism type 1B (PHP1B), also referred to as inactivating PTH/PTHrP signaling disorder (iPPSD), is characterized by proximal renal tubular resistance to parathyroid hormone (PTH) leading to hypocalcemia, hyperphosphatemia, and elevated PTH values. Autosomal dominant PHP1B (AD-PHP1B) with loss of methylation at the maternal GNAS A/B:TSS-DMR (transcription start site-differentially methylated region) alone can be caused by maternal deletions involving STX16.
Objective: Characterize a previously not reported AD-PHP1B family with loss of methylation at GNAS A/B:TSS-DMR, but without evidence for a STX16 deletion on the maternal allele and assess GNAS-AS2:TSS-DMR methylation.
Methods: DNA from 24 patients and 10 controls were investigated. AD-PHP1B patients without STX16 deletion from a single family (n = 5), AD-PHP1B patients with STX16 deletion (n = 9), sporPHP1B (n = 10), unaffected controls (n = 10), patUPD20 (n = 1), and matUPD20 (n = 1). Methylation and copy number analyses were performed by pyrosequencing, methylation-sensitive multiplex ligation-dependent probe amplification, and multiplex ligation-dependent probe amplification.
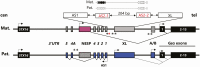
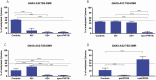
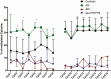
Results: Molecular cloning of polymerase chain reaction-amplified, bisulfite-treated genomic DNA from healthy controls revealed evidence for 2 distinct GNAS-AS2:TSS-DMR subdomains, named AS2-1 and AS2-2, which showed 16.0 ± 2.3% and 31.0 ± 2.2% methylation, respectively. DNA from affected members of a previously not reported AD-PHP1B family without the known genetic defects revealed incomplete loss of methylation at GNAS A/B:TSS-DMR, normal methylation at the 3 well-established maternal and paternal DMRs, and, surprisingly, increased methylation at AS2-1 (32.9 ± 3.5%), but not at AS2-2 (30.5 ± 2.9%).
Conclusion: The distinct methylation changes at the novel GNAS-AS2:TSS-DMR will help characterize further different PHP1B/iPPSD3 variants and will guide the search for underlying genetic defects, which may provide novel insights into the mechanisms underlying GNAS methylation.
Keywords: GNAS; DMR; iPPSD; imprinting; pseudohypoparathyroidism.
© The Author(s) 2021. Published by Oxford University Press on behalf of the Endocrine Society. All rights reserved. For permissions, please e-mail: journals.permissions@oup.com.
Figures

References
-
- Mantovani G. Clinical review: pseudohypoparathyroidism: diagnosis and treatment. J Clin Endocrinol Metab. 2011;96(10):3020-3030. - PubMed
-
- Weinstein LS, Yu S, Warner DR, Liu J. Endocrine manifestations of stimulatory G protein alpha-subunit mutations and the role of genomic imprinting. Endocr Rev. 2001;22(5):675-705. - PubMed
-
- Bastepe M, Jüppner H. Pseudohypoparathyroidism. New insights into an old disease. Endocrinol Metab Clin North Am. 2000;29(3):569-589. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

