Genetic Manipulation of Non-tuberculosis Mycobacteria
- PMID: 33679662
- PMCID: PMC7925387
- DOI: 10.3389/fmicb.2021.633510
Genetic Manipulation of Non-tuberculosis Mycobacteria
Abstract
Non-tuberculosis mycobacteria (NTMs) comprise a large group of organisms that are phenotypically diverse. Analysis of the growing number of completed NTM genomes has revealed both significant intra-genus genetic diversity, and a high percentage of predicted genes that appear to be unique to this group. Most NTMs have not been studied, however, the rise in NTM infections in several countries has prompted increasing interest in these organisms. Mycobacterial research has recently benefitted from the development of new genetic tools and a growing number of studies describing the genetic manipulation of NTMs have now been reported. In this review, we discuss the use of both site-specific and random mutagenesis tools in NTMs, highlighting the challenges that exist in applying these techniques to this diverse group of organisms.
Keywords: CRISPR/Cas; homologous recombination; mutagenesis; non-tuberculosis mycobacteria (NTMs); recombineering; transposon mutagenesis.
Copyright © 2021 Chimukuche and Williams.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

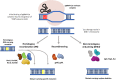
References
-
- Bardarov S., Bardarov S., Pavelka M. S., Sambandamurthy V., Larsen M., Tufariello J., et al. (2002). Specialized transduction: an efficient method for generating marked and unmarked targeted gene disruptions in Mycobacterium tuberculosis, M. bovis BCG and M. smegmatis. Microbiology 148 3007–3017. 10.1099/00221287-148-10-3007 - DOI - PubMed
-
- Budell W. C., Germain G. A., Janisch N., McKie−Krisberg Z., Jayaprakash A. D., Resnick A. E., et al. (2020). Transposon mutagenesis in Mycobacterium kansasii links a small RNA gene to colony morphology and biofilm formation and identifies 9,885 intragenic insertions that do not compromise colony outgrowth. MicrobiologyOpen 9:e988. 10.1002/mbo3.988 - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

