Comprehensive circRNA Expression Profile and Construction of circRNAs-Related ceRNA Network in a Mouse Model of Autism
- PMID: 33679870
- PMCID: PMC7928284
- DOI: 10.3389/fgene.2020.623584
Comprehensive circRNA Expression Profile and Construction of circRNAs-Related ceRNA Network in a Mouse Model of Autism
Abstract
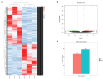
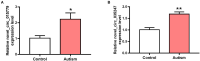
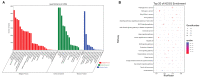
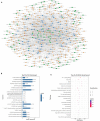
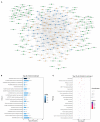
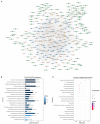
Autism is a common disease that seriously affects the quality of life. The role of circular RNAs (circRNAs) in autism remains largely unexplored. We aimed to detect the circRNA expression profile and construct a circRNA-based competing endogenous RNA (ceRNA) network in autism. Valproate acid was used to establish an in vivo model of autism in mice. A total of 1,059 differentially expressed circRNAs (477 upregulated and 582 downregulated) in autism group was identified by RNA sequencing. The expression of novel_circ_015779 and novel_circ_035247 were detected by real-time PCR. A ceRNA network based on altered circRNAs was established, with 9,715 nodes and 150,408 edges. Module analysis was conducted followed by GO and KEGG pathway enrichment analysis. The top three modules were all correlated with autism-related pathways involving "TGF-beta signaling pathway," "Notch signaling pathway," "MAPK signaling pathway," "long term depression," "thyroid hormone signaling pathway," etc. The present study reveals a novel circRNA involved mechanisms in the pathogenesis of autism.
Keywords: RNA sequencing (RNA-Seq); autism; ceRNA network; circular RNA (circRNA); in silico analysis.
Copyright © 2021 Wang, Yang, Chen, Xu, Wang, Liu, Zhang and Jiang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
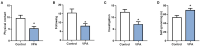

References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

