Loss of Ciliary Gene Bbs8 Results in Physiological Defects in the Retinal Pigment Epithelium
- PMID: 33681195
- PMCID: PMC7930748
- DOI: 10.3389/fcell.2021.607121
Loss of Ciliary Gene Bbs8 Results in Physiological Defects in the Retinal Pigment Epithelium
Abstract
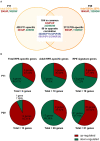
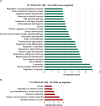
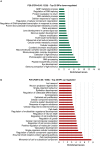
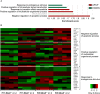
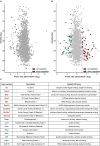
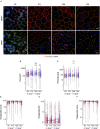
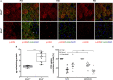
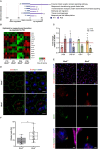
Primary cilia are sensory organelles vital for developmental and physiological processes. Their dysfunction causes a range of phenotypes including retinopathies. Although primary cilia have been described in the retinal pigment epithelium (RPE), little is known about their contribution to biological processes within this tissue. Ciliary proteins are increasingly being identified in non-ciliary locations and might carry out additional functions, disruption of which possibly contributes to pathology. The RPE is essential for maintaining photoreceptor cells and visual function. We demonstrate that upon loss of Bbs8, predominantly thought to be a ciliary gene, the RPE shows changes in gene and protein expression initially involved in signaling pathways and developmental processes, and at a later time point RPE homeostasis and function. Differentially regulated molecules affecting the cytoskeleton and cellular adhesion, led to defective cellular polarization and morphology associated with a possible epithelial-to-mesenchymal transition (EMT)-like phenotype. Our data highlights the benefit of combinatorial "omics" approaches with in vivo data for investigating the function of ciliopathy proteins. It also emphasizes the importance of ciliary proteins in the RPE and their contribution to visual disorders, which must be considered when designing treatment strategies for retinal degeneration.
Keywords: Epithelial-to-Mesenchym Transition (EMT); RPE; cilia; ciliopathy; genetic disease; molecular medicine; retinal pigment epithelium.
Copyright © 2021 Schneider, De Cegli, Nagarajan, Kretschmer, Matthiessen, Intartaglia, Hotaling, Ueffing, Boldt, Conte and May-Simera.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Primary Cilium-Mediated Retinal Pigment Epithelium Maturation Is Disrupted in Ciliopathy Patient Cells.Cell Rep. 2018 Jan 2;22(1):189-205. doi: 10.1016/j.celrep.2017.12.038. Cell Rep. 2018. PMID: 29298421 Free PMC article.
-
Deletion of IFT20 exclusively in the RPE ablates primary cilia and leads to retinal degeneration.PLoS Biol. 2023 Dec 4;21(12):e3002402. doi: 10.1371/journal.pbio.3002402. eCollection 2023 Dec. PLoS Biol. 2023. PMID: 38048369 Free PMC article.
-
Primary cilia in retinal pigment epithelium development and diseases.J Cell Mol Med. 2021 Oct;25(19):9084-9088. doi: 10.1111/jcmm.16882. Epub 2021 Aug 27. J Cell Mol Med. 2021. PMID: 34448530 Free PMC article. Review.
-
Cilia-associated wound repair mediated by IFT88 in retinal pigment epithelium.Sci Rep. 2023 May 21;13(1):8205. doi: 10.1038/s41598-023-35099-3. Sci Rep. 2023. PMID: 37211572 Free PMC article.
-
Role of Primary Cilia in the Eye.Adv Exp Med Biol. 2025;1468:441-445. doi: 10.1007/978-3-031-76550-6_72. Adv Exp Med Biol. 2025. PMID: 39930235 Review.
Cited by
-
The actin-bundling protein Fascin-1 modulates ciliary signalling.J Mol Cell Biol. 2023 Aug 3;15(4):mjad022. doi: 10.1093/jmcb/mjad022. J Mol Cell Biol. 2023. PMID: 37015875 Free PMC article.
-
Emerging principles of primary cilia dynamics in controlling tissue organization and function.EMBO J. 2023 Nov 2;42(21):e113891. doi: 10.15252/embj.2023113891. Epub 2023 Sep 25. EMBO J. 2023. PMID: 37743763 Free PMC article. Review.
-
TMEM97 governs partial epithelial-mesenchymal transition of retinal pigment epithelial cells via the CTNND2-ADAM10 axis.Mol Ther Nucleic Acids. 2025 Jan 21;36(1):102460. doi: 10.1016/j.omtn.2025.102460. eCollection 2025 Mar 11. Mol Ther Nucleic Acids. 2025. PMID: 39995975 Free PMC article.
-
Loss of Bbs8 leads to cystic kidney disease in mice with reduced acetylation of ciliary alpha-tubulin through HDAC2.bioRxiv [Preprint]. 2025 Jun 17:2024.03.07.583949. doi: 10.1101/2024.03.07.583949. bioRxiv. 2025. PMID: 40667253 Free PMC article. Preprint.
-
Intraflagellar transport: A critical player in photoreceptor development and the pathogenesis of retinal degenerative diseases.Cytoskeleton (Hoboken). 2024 Nov;81(11):556-568. doi: 10.1002/cm.21823. Epub 2023 Dec 23. Cytoskeleton (Hoboken). 2024. PMID: 38140908 Review.
References
-
- Adijanto J., Castorino J. J., Wang Z. X., Maminishkis A., Grunwald G. B., Philp N. J. (2012). Microphthalmia-associated transcription factor (MITF) promotes differentiation of human retinal pigment epithelium (RPE) by regulating microRNAs-204/211 expression. J. Biol. Chem. 287, 20491–20503. 10.1074/jbc.M112.354761 - DOI - PMC - PubMed
-
- Bates D., Mächler M., Bolker B., Walker S. (2015). Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 201–210. 10.18637/jss.v067.i01 - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

