Reproducing diabetic retinopathy features using newly developed human induced-pluripotent stem cell-derived retinal Müller glial cells
- PMID: 33683746
- PMCID: PMC8252429
- DOI: 10.1002/glia.23983
Reproducing diabetic retinopathy features using newly developed human induced-pluripotent stem cell-derived retinal Müller glial cells
Abstract
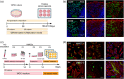
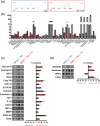
Muller glial cells (MGCs) are responsible for the homeostatic and metabolic support of the retina. Despite the importance of MGCs in retinal disorders, reliable and accessible human cell sources to be used to model MGC-associated diseases are lacking. Although primary human MGCs (pMGCs) can be purified from post-mortem retinal tissues, the donor scarcity limits their use. To overcome this problem, we developed a protocol to generate and bank human induced pluripotent stem cell-derived MGCs (hiMGCs). Using a transcriptome analysis, we showed that the three genetically independent hiMGCs generated were homogeneous and showed phenotypic characteristics and transcriptomic profile of pMGCs. These cells expressed key MGC markers, including Vimentin, CLU, DKK3, SOX9, SOX2, S100A16, ITGB1, and CD44 and could be cultured up to passage 8. Under our culture conditions, hiMGCs and pMGCs expressed low transcript levels of RLPB1, AQP4, KCNJ1, KCJN10, and SLC1A3. Using a disease modeling approach, we showed that hiMGCs could be used to model the features of diabetic retinopathy (DR)-associated dyslipidemia. Indeed, palmitate, a major free fatty acid with elevated plasma levels in diabetic patients, induced the expression of inflammatory cytokines found in the ocular fluid of DR patients such as CXCL8 (IL-8) and ANGPTL4. Moreover, the analysis of palmitate-treated hiMGC secretome showed an upregulation of proangiogenic factors strongly related to DR, including ANG2, Endoglin, IL-1β, CXCL8, MMP-9, PDGF-AA, and VEGF. Thus, hiMGCs could be an alternative to pMGCs and an extremely valuable tool to help to understand and model glial cell involvement in retinal disorders, including DR.
Keywords: Muller glial cells; diabetes; disease modeling; dyslipidemia; iPSC; retinopathy; stem cells.
© 2021 The Authors. GLIA published by Wiley Periodicals LLC.
Conflict of interest statement
OG, J‐AS, and SR are inventors on pending patents related to generation of retinal cells from human pluripotent stem cells.
Figures



References
-
- ACCORD Study group , ACCORD Eye Study Group , Chew, E. Y. , Ambrosius, W. T. , Davis, M. D. , Danis, R. P. , … Fine, L. J. (2010). Effects of medical therapies on retinopathy progression in type 2 diabetes. The New England Journal of Medicine, 363(3), 233–244. 10.1056/NEJMoa1001288 - DOI - PMC - PubMed
-
- Babapoor‐Farrokhran, S. , Jee, K. , Puchner, B. , Hassan, S. J. , Xin, X. , Rodrigues, M. , … Sodhi, A. (2015). Angiopoietin‐like 4 is a potent angiogenic factor and a novel therapeutic target for patients with proliferative diabetic retinopathy. Proceedings of the National Academy of Sciences of the United States of America, 112(23), E3030–E3039. 10.1073/pnas.1423765112 - DOI - PMC - PubMed
-
- Bromberg‐White, J. L. , Glazer, L. , Downer, R. , Furge, K. , Boguslawski, E. , & Duesbery, N. S. (2013). Identification of VEGF‐independent cytokines in proliferative diabetic retinopathy vitreous. Investigative Ophthalmology & Visual Science, 54(10), 6472–6480. 10.1167/iovs.13-12518 - DOI - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

