A flat embedding method for transmission electron microscopy reveals an unknown mechanism of tetracycline
- PMID: 33686188
- PMCID: PMC7940657
- DOI: 10.1038/s42003-021-01809-8
A flat embedding method for transmission electron microscopy reveals an unknown mechanism of tetracycline
Abstract
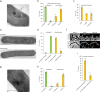
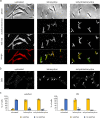
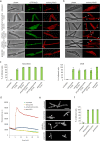
Transmission electron microscopy of cell sample sections is a popular technique in microbiology. Currently, ultrathin sectioning is done on resin-embedded cell pellets, which consumes milli- to deciliters of culture and results in sections of randomly orientated cells. This is problematic for rod-shaped bacteria and often precludes large-scale quantification of morphological phenotypes due to the lack of sufficient numbers of longitudinally cut cells. Here we report a flat embedding method that enables observation of thousands of longitudinally cut cells per single section and only requires microliter culture volumes. We successfully applied this technique to Bacillus subtilis, Escherichia coli, Mycobacterium bovis, and Acholeplasma laidlawii. To assess the potential of the technique to quantify morphological phenotypes, we monitored antibiotic-induced changes in B. subtilis cells. Surprisingly, we found that the ribosome inhibitor tetracycline causes membrane deformations. Further investigations showed that tetracycline disturbs membrane organization and localization of the peripheral membrane proteins MinD, MinC, and MreB. These observations are not the result of ribosome inhibition but constitute a secondary antibacterial activity of tetracycline that so far has defied discovery.
Conflict of interest statement
The authors declare that this work was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Systematic stress adaptation of Bacillus subtilis to tetracycline exposure.Ecotoxicol Environ Saf. 2020 Jan 30;188:109910. doi: 10.1016/j.ecoenv.2019.109910. Epub 2019 Nov 15. Ecotoxicol Environ Saf. 2020. PMID: 31740237
-
[Changes in the properties of the plasma membrane of Acholeplasma laidlawii under the action of tetracycline].Antibiot Khimioter. 1999;44(2):8-12. Antibiot Khimioter. 1999. PMID: 10202551 Russian.
-
Tetracycline inhibits propagation of deoxyribonucleic acid replication and alters membrane properties.Antimicrob Agents Chemother. 1977 Feb;11(2):318-23. doi: 10.1128/AAC.11.2.318. Antimicrob Agents Chemother. 1977. PMID: 403855 Free PMC article.
-
Functions of tetracycline efflux proteins that do not involve tetracycline.J Mol Microbiol Biotechnol. 2001 Apr;3(2):237-46. J Mol Microbiol Biotechnol. 2001. PMID: 11321579 Review.
-
Bacillus subtilis extracytoplasmic function (ECF) sigma factors and defense of the cell envelope.Curr Opin Microbiol. 2016 Apr;30:122-132. doi: 10.1016/j.mib.2016.02.002. Epub 2016 Feb 20. Curr Opin Microbiol. 2016. PMID: 26901131 Free PMC article. Review.
Cited by
-
Monitoring membranes: The exploration of biological bilayers with second harmonic generation.Chem Phys Rev. 2022 Dec;3(4):041307. doi: 10.1063/5.0120888. Epub 2022 Dec 14. Chem Phys Rev. 2022. PMID: 36536669 Free PMC article. Review.
-
Bacillus subtilis forms twisted cells with cell wall integrity defects upon removal of the molecular chaperones DnaK and trigger factor.Front Microbiol. 2023 Jan 16;13:988768. doi: 10.3389/fmicb.2022.988768. eCollection 2022. Front Microbiol. 2023. PMID: 36726573 Free PMC article.
-
Mechanistic insights into the C55-P targeting lipopeptide antibiotics revealed by structure-activity studies and high-resolution crystal structures.Chem Sci. 2022 Feb 21;13(10):2985-2991. doi: 10.1039/d1sc07190d. eCollection 2022 Mar 9. Chem Sci. 2022. PMID: 35382464 Free PMC article.
-
Mechanistic Studies and In Vivo Efficacy of an Oxadiazole-Containing Antibiotic.J Med Chem. 2022 May 12;65(9):6612-6630. doi: 10.1021/acs.jmedchem.1c02034. Epub 2022 Apr 28. J Med Chem. 2022. PMID: 35482444 Free PMC article.
-
Antimicrobial Activity of an Fmoc-Plantaricin 149 Derivative Peptide against Multidrug-Resistant Bacteria.Antibiotics (Basel). 2023 Feb 15;12(2):391. doi: 10.3390/antibiotics12020391. Antibiotics (Basel). 2023. PMID: 36830301 Free PMC article.
References
-
- Santhana Raj L. et al. Rapid method for transmission electron microscopy study of Staphylococcus aureus ATCC 25923. Ann. Microsc. 2007;7:102–108.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

